The title compound, 5-hydroxy-4′,7-dimethoxyisoflavone, C
17H
14O
5, is composed of a benzopyranone moiety, a phenyl moiety and two methoxy groups. The benzopyranone ring is not coplanar with the phenyl ring, the dihedral angle between them being 56.28 (3)°. The two methoxy groups are nearly coplanar with their corresponding rings, having C—C—O—C torsion angles of 2.9 (2) and 5.9 (2)°. The molecules are linked by C—H

O hydrogen bonds into sheets containing classical centrosymmetric
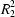
(8) rings. The sheets are further linked by aromatic π–π stacking interactions and C—H

O hydrogen bonds into a supramolecular structure.
Supporting information
CCDC reference: 223440
Genistein (1.0 g) was dissolved in Na2CO3 (20 ml, 5%) and dimethyl sulfate (0.5 ml) was added dropwise to the solution with stirring. The mixture was stirred for 4 h at room temperature and a colorless precipitate began to appear. The precipitate was filtered off and washed with water until the pH of the filtrate was 8. After recrystallization from ethyl acetate, the product had a melting point of 428 K. Crystals of (I) suitable for X-ray analysis were obtained by slow evaporation from ethyl acetate after 7 d at room temperature.
H atoms were placed at calculated positions and treated as riding, with C—H distances in the range 0.93–0.96 Å and Uiso(H) values of 1.2Ueq of the attached C atom [1.5Ueq(C) for methyl H atoms].
Data collection: XSCANS (Siemens, 1996); cell refinement: XSCANS; data reduction: SHELXTL (Siemens, 1997); program(s) used to solve structure: SHELXS97 (Sheldrick, 1990); program(s) used to refine structure: SHELXL97 (Sheldrick, 1997); molecular graphics: SHELXTL; software used to prepare material for publication: SHELXTL.
5-hydroxy-4',7-dimethoxyisoflavone
top
Crystal data top
| C17H14O5 | F(000) = 624 |
| Mr = 298.28 | Dx = 1.441 Mg m−3 |
| Monoclinic, P21/n | Melting point: 428 K |
| Hall symbol: -P 2yn | Mo Kα radiation, λ = 0.71073 Å |
| a = 5.7754 (8) Å | Cell parameters from 34 reflections |
| b = 7.9446 (12) Å | θ = 2.7–15.9° |
| c = 30.044 (5) Å | µ = 0.11 mm−1 |
| β = 93.807 (12)° | T = 296 K |
| V = 1375.4 (4) Å3 | Prism, colorless |
| Z = 4 | 0.58 × 0.54 × 0.50 mm |
Data collection top
Siemens P4
diffractometer | Rint = 0.010 |
| Radiation source: normal-focus sealed tube | θmax = 25.3°, θmin = 1.4° |
| Graphite monochromator | h = 0→6 |
| ω scans | k = 0→9 |
| 3089 measured reflections | l = −36→36 |
| 2485 independent reflections | 3 standard reflections every 97 reflections |
| 1673 reflections with I > 2σ(I) | intensity decay: 0.4% |
Refinement top
| Refinement on F2 | Secondary atom site location: difference Fourier map |
| Least-squares matrix: full | Hydrogen site location: inferred from neighbouring sites |
| R[F2 > 2σ(F2)] = 0.035 | H-atom parameters constrained |
| wR(F2) = 0.087 | w = 1/[σ2(Fo2) + (0.0522P)2]
where P = (Fo2 + 2Fc2)/3 |
| S = 0.90 | (Δ/σ)max = 0.001 |
| 2485 reflections | Δρmax = 0.19 e Å−3 |
| 203 parameters | Δρmin = −0.15 e Å−3 |
| 0 restraints | Extinction correction: SHELXL97, Fc*=kFc[1+0.001xFc2λ3/sin(2θ)]-1/4 |
| Primary atom site location: structure-invariant direct methods | Extinction coefficient: 0.0233 (18) |
Crystal data top
| C17H14O5 | V = 1375.4 (4) Å3 |
| Mr = 298.28 | Z = 4 |
| Monoclinic, P21/n | Mo Kα radiation |
| a = 5.7754 (8) Å | µ = 0.11 mm−1 |
| b = 7.9446 (12) Å | T = 296 K |
| c = 30.044 (5) Å | 0.58 × 0.54 × 0.50 mm |
| β = 93.807 (12)° | |
Data collection top
Siemens P4
diffractometer | Rint = 0.010 |
| 3089 measured reflections | 3 standard reflections every 97 reflections |
| 2485 independent reflections | intensity decay: 0.4% |
| 1673 reflections with I > 2σ(I) | |
Refinement top
| R[F2 > 2σ(F2)] = 0.035 | 0 restraints |
| wR(F2) = 0.087 | H-atom parameters constrained |
| S = 0.90 | Δρmax = 0.19 e Å−3 |
| 2485 reflections | Δρmin = −0.15 e Å−3 |
| 203 parameters | |
Special details top
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes. |
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2, conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2 are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| O1 | −0.01497 (17) | 0.23550 (14) | 0.65150 (3) | 0.0479 (3) | |
| O2 | 0.67377 (18) | 0.55119 (15) | 0.62862 (4) | 0.0529 (3) | |
| H2O | 0.6810 | 0.5506 | 0.6560 | 0.064* | |
| O3 | 0.54957 (17) | 0.48501 (15) | 0.70802 (3) | 0.0501 (3) | |
| O4 | 0.2227 (2) | 0.34532 (17) | 0.50583 (4) | 0.0652 (4) | |
| O5 | 0.2007 (2) | 0.38703 (16) | 0.90489 (3) | 0.0589 (3) | |
| C1 | 0.1410 (3) | 0.30694 (19) | 0.62475 (5) | 0.0407 (4) | |
| C2 | 0.0933 (3) | 0.2862 (2) | 0.57932 (5) | 0.0470 (4) | |
| H2 | −0.0376 | 0.2279 | 0.5681 | 0.056* | |
| C3 | 0.2472 (3) | 0.3554 (2) | 0.55126 (5) | 0.0476 (4) | |
| C4 | 0.4436 (3) | 0.4439 (2) | 0.56772 (5) | 0.0484 (4) | |
| H4 | 0.5449 | 0.4897 | 0.5482 | 0.058* | |
| C5 | 0.4862 (2) | 0.46291 (19) | 0.61293 (5) | 0.0411 (4) | |
| C6 | 0.3343 (2) | 0.39272 (18) | 0.64309 (5) | 0.0377 (4) | |
| C7 | 0.3740 (2) | 0.41124 (19) | 0.69099 (5) | 0.0388 (4) | |
| C8 | 0.1961 (2) | 0.34029 (19) | 0.71724 (5) | 0.0392 (4) | |
| C9 | 0.0199 (3) | 0.2566 (2) | 0.69617 (5) | 0.0460 (4) | |
| H9 | −0.0887 | 0.2085 | 0.7139 | 0.055* | |
| C10 | 0.2007 (2) | 0.35938 (19) | 0.76658 (5) | 0.0399 (4) | |
| C11 | 0.0120 (3) | 0.4273 (2) | 0.78600 (5) | 0.0446 (4) | |
| H11 | −0.1137 | 0.4652 | 0.7678 | 0.054* | |
| C12 | 0.0040 (3) | 0.4406 (2) | 0.83202 (5) | 0.0455 (4) | |
| H12 | −0.1245 | 0.4876 | 0.8444 | 0.055* | |
| C13 | 0.1896 (3) | 0.3833 (2) | 0.85900 (5) | 0.0430 (4) | |
| C14 | 0.3817 (3) | 0.3153 (2) | 0.84024 (5) | 0.0474 (4) | |
| H14 | 0.5065 | 0.2762 | 0.8585 | 0.057* | |
| C15 | 0.3877 (3) | 0.3056 (2) | 0.79459 (5) | 0.0447 (4) | |
| H15 | 0.5188 | 0.2624 | 0.7823 | 0.054* | |
| C16 | 0.0224 (3) | 0.2657 (2) | 0.48580 (6) | 0.0693 (6) | |
| H16A | 0.0271 | 0.1478 | 0.4927 | 0.083* | |
| H16B | 0.0183 | 0.2805 | 0.4540 | 0.083* | |
| H16C | −0.1138 | 0.3149 | 0.4971 | 0.083* | |
| C17 | −0.0006 (3) | 0.4395 (3) | 0.92563 (6) | 0.0701 (6) | |
| H17A | −0.0329 | 0.5551 | 0.9183 | 0.084* | |
| H17B | 0.0246 | 0.4278 | 0.9574 | 0.084* | |
| H17C | −0.1300 | 0.3710 | 0.9152 | 0.084* | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| O1 | 0.0451 (6) | 0.0544 (7) | 0.0445 (6) | −0.0130 (5) | 0.0053 (5) | 0.0019 (5) |
| O2 | 0.0459 (6) | 0.0624 (8) | 0.0509 (7) | −0.0150 (6) | 0.0062 (5) | 0.0014 (6) |
| O3 | 0.0384 (6) | 0.0637 (8) | 0.0478 (6) | −0.0080 (6) | −0.0006 (5) | 0.0031 (6) |
| O4 | 0.0744 (8) | 0.0792 (9) | 0.0421 (7) | −0.0131 (7) | 0.0041 (6) | −0.0062 (6) |
| O5 | 0.0617 (7) | 0.0739 (9) | 0.0410 (6) | 0.0041 (7) | 0.0022 (6) | 0.0013 (6) |
| C1 | 0.0385 (8) | 0.0380 (9) | 0.0462 (9) | −0.0022 (7) | 0.0068 (7) | 0.0040 (7) |
| C2 | 0.0459 (9) | 0.0469 (10) | 0.0479 (9) | −0.0066 (8) | −0.0002 (7) | −0.0031 (8) |
| C3 | 0.0550 (10) | 0.0488 (10) | 0.0395 (9) | 0.0025 (9) | 0.0058 (8) | −0.0005 (7) |
| C4 | 0.0485 (10) | 0.0514 (10) | 0.0465 (10) | −0.0047 (8) | 0.0119 (8) | 0.0052 (8) |
| C5 | 0.0361 (8) | 0.0397 (9) | 0.0482 (9) | −0.0004 (7) | 0.0068 (7) | 0.0016 (7) |
| C6 | 0.0351 (8) | 0.0362 (8) | 0.0420 (9) | 0.0024 (7) | 0.0048 (6) | 0.0036 (7) |
| C7 | 0.0327 (8) | 0.0383 (9) | 0.0452 (9) | 0.0037 (7) | 0.0011 (7) | 0.0035 (7) |
| C8 | 0.0344 (8) | 0.0392 (9) | 0.0440 (8) | 0.0023 (7) | 0.0034 (7) | 0.0059 (7) |
| C9 | 0.0426 (9) | 0.0497 (10) | 0.0465 (9) | −0.0048 (8) | 0.0094 (7) | 0.0063 (8) |
| C10 | 0.0359 (8) | 0.0404 (9) | 0.0435 (9) | −0.0013 (7) | 0.0046 (7) | 0.0059 (7) |
| C11 | 0.0362 (8) | 0.0520 (10) | 0.0456 (9) | 0.0035 (7) | 0.0020 (7) | 0.0100 (8) |
| C12 | 0.0385 (8) | 0.0524 (10) | 0.0463 (9) | 0.0053 (8) | 0.0085 (7) | 0.0052 (8) |
| C13 | 0.0442 (9) | 0.0450 (9) | 0.0398 (9) | −0.0045 (8) | 0.0022 (7) | 0.0034 (7) |
| C14 | 0.0395 (9) | 0.0529 (10) | 0.0485 (10) | 0.0051 (8) | −0.0060 (7) | 0.0055 (8) |
| C15 | 0.0362 (8) | 0.0493 (10) | 0.0488 (10) | 0.0040 (7) | 0.0045 (7) | 0.0033 (8) |
| C16 | 0.0804 (13) | 0.0746 (14) | 0.0512 (11) | −0.0036 (12) | −0.0076 (10) | −0.0113 (10) |
| C17 | 0.0696 (12) | 0.0936 (16) | 0.0482 (10) | −0.0055 (11) | 0.0125 (9) | −0.0068 (10) |
Geometric parameters (Å, º) top
| O1—C9 | 1.3542 (17) | C8—C9 | 1.339 (2) |
| O1—C1 | 1.3697 (17) | C8—C10 | 1.489 (2) |
| O2—C5 | 1.3489 (17) | C9—H9 | 0.9300 |
| O2—H2O | 0.8200 | C10—C11 | 1.380 (2) |
| O3—C7 | 1.2508 (17) | C10—C15 | 1.3913 (19) |
| O4—C3 | 1.3653 (18) | C11—C12 | 1.390 (2) |
| O4—C16 | 1.4167 (19) | C11—H11 | 0.9300 |
| O5—C13 | 1.3762 (18) | C12—C13 | 1.378 (2) |
| O5—C17 | 1.418 (2) | C12—H12 | 0.9300 |
| C1—C2 | 1.384 (2) | C13—C14 | 1.387 (2) |
| C1—C6 | 1.390 (2) | C14—C15 | 1.377 (2) |
| C2—C3 | 1.379 (2) | C14—H14 | 0.9300 |
| C2—H2 | 0.9300 | C15—H15 | 0.9300 |
| C3—C4 | 1.397 (2) | C16—H16A | 0.9600 |
| C4—C5 | 1.373 (2) | C16—H16B | 0.9600 |
| C4—H4 | 0.9300 | C16—H16C | 0.9600 |
| C5—C6 | 1.417 (2) | C17—H17A | 0.9600 |
| C6—C7 | 1.4497 (19) | C17—H17B | 0.9600 |
| C7—C8 | 1.450 (2) | C17—H17C | 0.9600 |
| | | |
| C9—O1—C1 | 118.05 (12) | C11—C10—C15 | 117.87 (13) |
| C5—O2—H2O | 109.5 | C11—C10—C8 | 119.89 (13) |
| C3—O4—C16 | 118.58 (13) | C15—C10—C8 | 122.20 (13) |
| C13—O5—C17 | 117.49 (13) | C10—C11—C12 | 121.97 (14) |
| O1—C1—C2 | 115.71 (13) | C10—C11—H11 | 119.0 |
| O1—C1—C6 | 120.83 (13) | C12—C11—H11 | 119.0 |
| C2—C1—C6 | 123.46 (14) | C13—C12—C11 | 118.95 (15) |
| C3—C2—C1 | 117.46 (15) | C13—C12—H12 | 120.5 |
| C3—C2—H2 | 121.3 | C11—C12—H12 | 120.5 |
| C1—C2—H2 | 121.3 | O5—C13—C12 | 124.41 (14) |
| O4—C3—C2 | 124.01 (15) | O5—C13—C14 | 115.48 (13) |
| O4—C3—C4 | 114.29 (14) | C12—C13—C14 | 120.11 (14) |
| C2—C3—C4 | 121.70 (14) | C15—C14—C13 | 120.04 (14) |
| C5—C4—C3 | 119.58 (15) | C15—C14—H14 | 120.0 |
| C5—C4—H4 | 120.2 | C13—C14—H14 | 120.0 |
| C3—C4—H4 | 120.2 | C14—C15—C10 | 121.03 (14) |
| O2—C5—C4 | 119.34 (14) | C14—C15—H15 | 119.5 |
| O2—C5—C6 | 119.88 (13) | C10—C15—H15 | 119.5 |
| C4—C5—C6 | 120.78 (14) | O4—C16—H16A | 109.5 |
| C1—C6—C5 | 117.03 (13) | O4—C16—H16B | 109.5 |
| C1—C6—C7 | 120.89 (13) | H16A—C16—H16B | 109.5 |
| C5—C6—C7 | 122.08 (13) | O4—C16—H16C | 109.5 |
| O3—C7—C6 | 121.61 (14) | H16A—C16—H16C | 109.5 |
| O3—C7—C8 | 122.96 (14) | H16B—C16—H16C | 109.5 |
| C6—C7—C8 | 115.42 (13) | O5—C17—H17A | 109.5 |
| C9—C8—C7 | 118.66 (14) | O5—C17—H17B | 109.5 |
| C9—C8—C10 | 118.96 (13) | H17A—C17—H17B | 109.5 |
| C7—C8—C10 | 122.37 (13) | O5—C17—H17C | 109.5 |
| C8—C9—O1 | 126.03 (14) | H17A—C17—H17C | 109.5 |
| C8—C9—H9 | 117.0 | H17B—C17—H17C | 109.5 |
| O1—C9—H9 | 117.0 | | |
| | | |
| C9—O1—C1—C2 | −177.98 (14) | O3—C7—C8—C9 | −176.87 (14) |
| C9—O1—C1—C6 | 2.5 (2) | C6—C7—C8—C9 | 3.7 (2) |
| O1—C1—C2—C3 | −179.58 (14) | O3—C7—C8—C10 | 4.0 (2) |
| C6—C1—C2—C3 | −0.1 (2) | C6—C7—C8—C10 | −175.34 (13) |
| C16—O4—C3—C2 | 2.9 (2) | C7—C8—C9—O1 | −2.5 (2) |
| C16—O4—C3—C4 | −176.73 (15) | C10—C8—C9—O1 | 176.65 (14) |
| C1—C2—C3—O4 | −179.93 (15) | C1—O1—C9—C8 | −0.8 (2) |
| C1—C2—C3—C4 | −0.3 (2) | C9—C8—C10—C11 | −54.1 (2) |
| O4—C3—C4—C5 | 179.78 (14) | C7—C8—C10—C11 | 124.94 (16) |
| C2—C3—C4—C5 | 0.1 (2) | C9—C8—C10—C15 | 123.66 (17) |
| C3—C4—C5—O2 | −178.75 (14) | C7—C8—C10—C15 | −57.3 (2) |
| C3—C4—C5—C6 | 0.5 (2) | C15—C10—C11—C12 | −0.8 (2) |
| O1—C1—C6—C5 | −179.89 (13) | C8—C10—C11—C12 | 177.07 (15) |
| C2—C1—C6—C5 | 0.7 (2) | C10—C11—C12—C13 | −0.5 (2) |
| O1—C1—C6—C7 | −1.0 (2) | C17—O5—C13—C12 | 5.9 (2) |
| C2—C1—C6—C7 | 179.56 (14) | C17—O5—C13—C14 | −173.76 (16) |
| O2—C5—C6—C1 | 178.37 (13) | C11—C12—C13—O5 | −178.89 (14) |
| C4—C5—C6—C1 | −0.9 (2) | C11—C12—C13—C14 | 0.8 (2) |
| O2—C5—C6—C7 | −0.5 (2) | O5—C13—C14—C15 | 179.94 (15) |
| C4—C5—C6—C7 | −179.72 (14) | C12—C13—C14—C15 | 0.2 (2) |
| C1—C6—C7—O3 | 178.48 (14) | C13—C14—C15—C10 | −1.6 (2) |
| C5—C6—C7—O3 | −2.7 (2) | C11—C10—C15—C14 | 1.9 (2) |
| C1—C6—C7—C8 | −2.1 (2) | C8—C10—C15—C14 | −175.98 (15) |
| C5—C6—C7—C8 | 176.70 (14) | | |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| O2—H2O···O3 | 0.82 | 1.86 | 2.590 (2) | 148 |
| C4—H4···O4i | 0.93 | 2.54 | 3.459 (2) | 171 |
| C11—H11···O3ii | 0.93 | 2.56 | 3.464 (2) | 164 |
| C14—H14···O2iii | 0.93 | 2.58 | 3.399 (2) | 147 |
| Symmetry codes: (i) −x+1, −y+1, −z+1; (ii) x−1, y, z; (iii) −x+3/2, y−1/2, −z+3/2. |
Experimental details
| Crystal data |
| Chemical formula | C17H14O5 |
| Mr | 298.28 |
| Crystal system, space group | Monoclinic, P21/n |
| Temperature (K) | 296 |
| a, b, c (Å) | 5.7754 (8), 7.9446 (12), 30.044 (5) |
| β (°) | 93.807 (12) |
| V (Å3) | 1375.4 (4) |
| Z | 4 |
| Radiation type | Mo Kα |
| µ (mm−1) | 0.11 |
| Crystal size (mm) | 0.58 × 0.54 × 0.50 |
| |
| Data collection |
| Diffractometer | Siemens P4
diffractometer |
| Absorption correction | – |
No. of measured, independent and
observed [I > 2σ(I)] reflections | 3089, 2485, 1673 |
| Rint | 0.010 |
| (sin θ/λ)max (Å−1) | 0.600 |
| |
| Refinement |
| R[F2 > 2σ(F2)], wR(F2), S | 0.035, 0.087, 0.90 |
| No. of reflections | 2485 |
| No. of parameters | 203 |
| H-atom treatment | H-atom parameters constrained |
| Δρmax, Δρmin (e Å−3) | 0.19, −0.15 |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| O2—H2O···O3 | 0.82 | 1.86 | 2.590 (2) | 148 |
| C4—H4···O4i | 0.93 | 2.54 | 3.459 (2) | 171 |
| C11—H11···O3ii | 0.93 | 2.56 | 3.464 (2) | 164 |
| C14—H14···O2iii | 0.93 | 2.58 | 3.399 (2) | 147 |
| Symmetry codes: (i) −x+1, −y+1, −z+1; (ii) x−1, y, z; (iii) −x+3/2, y−1/2, −z+3/2. |
 O hydrogen bonds into sheets containing classical centrosymmetric
O hydrogen bonds into sheets containing classical centrosymmetric  O hydrogen bonds into a supramolecular structure.
O hydrogen bonds into a supramolecular structure.
 journal menu
journal menu








![[Figure 1]](http://journals.iucr.org/c/issues/2005/01/00/fr1473/fr1473fig1thm.gif)
![[Figure 2]](http://journals.iucr.org/c/issues/2005/01/00/fr1473/fr1473fig2thm.gif)
![[Figure 3]](http://journals.iucr.org/c/issues/2005/01/00/fr1473/fr1473fig3thm.gif)




Hydrogen bonds and π–π stacking interactions are an important research area in supramolecular chemistry and crystal engineering (MacDonald & Whitesides, 1994). These interactions play an important role in self-assembly and recognition of aromatic compounds (Janiak, 2000; Hunter & Sanders, 1990) as an auxiliary stabilizing short contact (William et al., 1999; Luque et al., 2001; Kaafarani et al., 2001). In biomacromolecular systems, stacking interactions and hydrogen bonds are important for the double-helical DNA structure (Hunter, 1993); they can direct the intercalation of drugs into DNA (Wang et al., 1984) and they contribute to the stability of the tertiary structure of proteins (Burley & Petsko, 1985).
Genistein, a natural soy isoflavone, has potential phytoestrogen (Hua et al., 2003; Warren, 2002) and antioxidant activities (Ian et al., 1995). Studies have also found genistein effective in inhibiting cardiovascular disease (Hwang et al., 2001), tyrosine kinases (Nevala et al., 2002), and cancer cell growth (Yuan et al., 2003; W. F. Chen et al., 2003), and in accelerating the formation of bone cells (X. W. Chen et al., 2003). The title compound, 5-hydroxy-4',7-dimethoxyisoflavone, (I), is a derivative of genistein, with potential medical applications. We report here the crystal structure of (I).
The title compound is composed of a benzopyranone moiety, a phenyl moiety and two methoxy groups (Fig. 1). The geometry of the isoflavone skeleton of (I) is similar to its analogue dalspinin (Lakshmi, et al., 1996) with respect to most of the bond distances and angles. The atoms of the benzopyranone moiety, including rings A (C1–C6) and C (O1/C1/C6–C9), are almost coplanar, the dihedral angle between ring A and ring C being 1.37 (8)°. To avoid steric conflicts, the two rigid ring systems, viz. phenyl ring B (C10–C15) and the benzopyranone moiety, are rotated by 56.28 (3)° with respect to one another. The methoxy group at atom C3 is nearly coplanar with ring A, as indicated by the C16—O4—C3—C2 torsion angle [2.9 (2)°]; the methoxy group at atom C13 is also coplanar with the attached ring, the C17—O5—C13—C12 torsion angle being 5.9 (2)°.
Fig. 2 shows how a cyclic dimer is formed through a supramolecular synthon, R22(8). Methoxy atom O4 acts as a hydrogen-bond acceptor, via atom H4, to atom C4 of ring A. In this manner, a centrosymmetric R22(8) ring is formed. Hydroxy atoms O2 from the two molecules linked by the R22(8) ring act as hydrogen-bond acceptors, via atoms H14, to atoms C14 of rings B in adjacent molecules. The combination of the C14—H14···O2 interaction and the R22(8) supramolecular synthon generates a (10=1) sheet, which includes two A, two B and two C rings from four molecules, and these six rings are almost coplanar; furthermore, these dimers are also linked into (100) chains by C11—H11···O3 interactions (Fig. 3). The combination of the (10=1) sheets and the (100) chains generates a three-dimensional framework. An independent O2—H2O···O3 intramolecular hydrogen bonds generates a characteristic intramolecular S(6) motif. Details of the hydrogen bonding are given in Table 1.
Intermolecular stacking via aromatic π–π interactions is also present (Fig. 3), the two molecules being offset by partial overlap of rings B (π rich) and C (π deficient). Ring B of one molecule and ring C of a neighbouring molecule are almost parallel, with a dihedral angle between them of 8.02 (7)°. The perpendicular plane-to-plane distance between the rings is 3.311 Å, and the corresponding Cg···Cgi distance is 3.693 Å [Cg represents the centroids of rings A and C; symmetry code: (i) 0.5 − x, 0.5 + y, 1.5 − z], indicating that a strong π–π stacking interaction exists in the title compound. Hydrogen bonds and aromatic π–π stacking interactions play a key role in assembling the supramolecular structure.