Buy article online - an online subscription or single-article purchase is required to access this article.
The title compound, C
4H
4N
6O
4·C
2H
6OS, crystallizes in the triclinic space group
P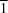
. It is a dimethyl sulfoxide (DMSO) solvate of the insensitive energetic compound 2,6-diamino-3,5-dinitro-1,4-pyrazine (ANPZ). The structure has been determined to obtain more accurate metrical parameters for the 2,6-diamino-3,5-dinitro-1,4-pyrazine moiety than those obtained from unsolvated crystals which are invariably twinned. The packing motif consists of two formula units linked by strong complementary hydrogen bonds between the ANPZ units with the two DMSO solvate molecules each linked by two bifurcated hydogen bonds to the two ANPZ units. This central motif is further linked into planar sheets by weaker interactions between the DMSO methyl H atoms and the O atoms from the nitro groups. This looser packing arrangement compared to ANPZ is reflected in a lower density (1.662 Mg m
-3 versus 1.812 Mg m
-3 for ANPZ).
Supporting information
CCDC reference: 170918
Key indicators
- Single-crystal X-ray study
- T = 93 K
- Mean
![[sigma]](//journals.iucr.org/logos/entities/sigma_rmgif.gif) (C-C) = 0.002 Å
(C-C) = 0.002 Å
- R factor = 0.035
- wR factor = 0.091
- Data-to-parameter ratio = 12.9
checkCIF results
No syntax errors found
ADDSYM reports no extra symmetry
 Alert Level C:
Alert Level C:
PLAT_030 Alert C Refined Extinction parameter within range .... 2.50 Sigma
0 Alert Level A = Potentially serious problem
0 Alert Level B = Potential problem
1 Alert Level C = Please check
Crystals of the title compound were supplied by Dr Philip Pagoria, Energetic
Materials Laboratory, Lawrence Livermore National Laboratory, Livermore, CA
94550, USA. Crystal and reflection data were obtained using standard
procedures (Butcher et al., 1995).
Data collection: SMART (Bruker, 1994); cell refinement: SMART; data reduction: SHELXTL (Bruker, 1994); program(s) used to solve structure: SHELXS97 (Sheldrick, 1990); program(s) used to refine structure: SHELXL97 (Sheldrick, 1997); molecular graphics: SHELXTL; software used to prepare material for publication: SHELXTL.
2,6-diamino-3,5-dinitro-1,4-pyrazine dimethyl sulfoxide solvate
top
Crystal data top
| C4H4N6O4·C2H6OS | Z = 2 |
| Mr = 278.26 | F(000) = 288 |
| Triclinic, P1 | Dx = 1.662 Mg m−3 |
| a = 5.7817 (5) Å | Mo Kα radiation, λ = 0.71073 Å |
| b = 8.1353 (8) Å | Cell parameters from 3759 reflections |
| c = 12.0270 (11) Å | θ = 2.5–28.3° |
| α = 99.253 (2)° | µ = 0.32 mm−1 |
| β = 94.113 (2)° | T = 93 K |
| γ = 92.482 (2)° | Prism, yellow |
| V = 556.04 (9) Å3 | 0.65 × 0.22 × 0.07 mm |
Data collection top
CCD area detector
diffractometer | 2638 independent reflections |
| Radiation source: fine-focus sealed tube | 2505 reflections with I > 2σ(I) |
| Graphite monochromator | Rint = 0.031 |
| ϕ and ω scans | θmax = 28.3°, θmin = 1.7° |
Absorption correction: integration
(Wuensch & Prewitt, 1965) | h = −6→7 |
| Tmin = 0.919, Tmax = 0.969 | k = −10→10 |
| 4175 measured reflections | l = −15→16 |
Refinement top
| Refinement on F2 | Secondary atom site location: difference Fourier map |
| Least-squares matrix: full | Hydrogen site location: inferred from neighbouring sites |
| R[F2 > 2σ(F2)] = 0.035 | H atoms treated by a mixture of independent and constrained refinement |
| wR(F2) = 0.091 | w = 1/[σ2(Fo2) + (0.0434P)2 + 0.3143P]
where P = (Fo2 + 2Fc2)/3 |
| S = 1.14 | (Δ/σ)max = 0.006 |
| 2638 reflections | Δρmax = 0.44 e Å−3 |
| 204 parameters | Δρmin = −0.25 e Å−3 |
| 0 restraints | Extinction correction: SHELXL97, Fc*=kFc[1+0.001xFc2λ3/sin(2θ)]-1/4 |
| Primary atom site location: structure-invariant direct methods | Extinction coefficient: 0.005 (2) |
Crystal data top
| C4H4N6O4·C2H6OS | γ = 92.482 (2)° |
| Mr = 278.26 | V = 556.04 (9) Å3 |
| Triclinic, P1 | Z = 2 |
| a = 5.7817 (5) Å | Mo Kα radiation |
| b = 8.1353 (8) Å | µ = 0.32 mm−1 |
| c = 12.0270 (11) Å | T = 93 K |
| α = 99.253 (2)° | 0.65 × 0.22 × 0.07 mm |
| β = 94.113 (2)° | |
Data collection top
CCD area detector
diffractometer | 2638 independent reflections |
Absorption correction: integration
(Wuensch & Prewitt, 1965) | 2505 reflections with I > 2σ(I) |
| Tmin = 0.919, Tmax = 0.969 | Rint = 0.031 |
| 4175 measured reflections | |
Refinement top
| R[F2 > 2σ(F2)] = 0.035 | 0 restraints |
| wR(F2) = 0.091 | H atoms treated by a mixture of independent and constrained refinement |
| S = 1.14 | Δρmax = 0.44 e Å−3 |
| 2638 reflections | Δρmin = −0.25 e Å−3 |
| 204 parameters | |
Special details top
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes)
are estimated using the full covariance matrix. The cell e.s.d.'s are taken
into account individually in the estimation of e.s.d.'s in distances, angles
and torsion angles; correlations between e.s.d.'s in cell parameters are only
used when they are defined by crystal symmetry. An approximate (isotropic)
treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s.
planes. |
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor
wR and goodness of fit S are based on F2, conventional
R-factors R are based on F, with F set to zero for
negative F2. The threshold expression of F2 >
σ(F2) is used only for calculating R-factors(gt) etc.
and is not relevant to the choice of reflections for refinement.
R-factors based on F2 are statistically about twice as large
as those based on F, and R- factors based on ALL data will be
even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| S1S | 1.17993 (6) | 0.66666 (4) | 1.34974 (3) | 0.01204 (11) | |
| O1S | 1.03544 (19) | 0.62121 (14) | 1.23767 (9) | 0.0172 (2) | |
| N1 | 0.7338 (2) | 0.64907 (15) | 0.94637 (10) | 0.0127 (2) | |
| C2 | 0.5825 (2) | 0.61920 (17) | 0.85410 (12) | 0.0123 (3) | |
| N2 | 0.3974 (2) | 0.51846 (17) | 0.85675 (11) | 0.0159 (3) | |
| H2A | 0.299 (4) | 0.494 (2) | 0.8020 (18) | 0.016 (5)* | |
| H2B | 0.379 (4) | 0.480 (3) | 0.9132 (19) | 0.017 (5)* | |
| C3 | 0.6322 (2) | 0.69773 (17) | 0.75867 (12) | 0.0121 (3) | |
| N3 | 0.4786 (2) | 0.67906 (15) | 0.65560 (10) | 0.0134 (2) | |
| O3A | 0.29497 (19) | 0.59424 (15) | 0.65159 (10) | 0.0215 (3) | |
| O3B | 0.53576 (19) | 0.74861 (15) | 0.57813 (9) | 0.0193 (2) | |
| N4 | 0.8207 (2) | 0.79105 (15) | 0.75808 (10) | 0.0123 (2) | |
| C5 | 0.9692 (2) | 0.81529 (17) | 0.84709 (12) | 0.0123 (3) | |
| N5 | 1.1745 (2) | 0.91916 (15) | 0.83599 (11) | 0.0140 (2) | |
| O5A | 1.32413 (18) | 0.94817 (14) | 0.91680 (10) | 0.0190 (2) | |
| O5B | 1.1932 (2) | 0.97274 (15) | 0.74713 (10) | 0.0208 (2) | |
| C6 | 0.9299 (2) | 0.74434 (17) | 0.94705 (12) | 0.0121 (3) | |
| N6 | 1.0750 (2) | 0.76880 (17) | 1.03955 (11) | 0.0155 (3) | |
| H6A | 1.207 (4) | 0.820 (3) | 1.0382 (17) | 0.020 (5)* | |
| H6B | 1.048 (4) | 0.724 (3) | 1.094 (2) | 0.023 (5)* | |
| C1S | 0.9899 (3) | 0.7636 (2) | 1.44803 (14) | 0.0205 (3) | |
| H1SA | 0.877 (4) | 0.682 (3) | 1.458 (2) | 0.038 (6)* | |
| H1SB | 0.920 (4) | 0.848 (3) | 1.4155 (19) | 0.026 (5)* | |
| H1SC | 1.080 (4) | 0.803 (3) | 1.518 (2) | 0.031 (6)* | |
| C2S | 1.3558 (3) | 0.8471 (2) | 1.33623 (15) | 0.0215 (3) | |
| H2SA | 1.463 (5) | 0.817 (3) | 1.282 (2) | 0.041 (7)* | |
| H2SB | 1.441 (4) | 0.884 (3) | 1.407 (2) | 0.029 (6)* | |
| H2SC | 1.259 (4) | 0.932 (3) | 1.315 (2) | 0.037 (6)* | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| S1S | 0.01286 (18) | 0.01223 (18) | 0.01090 (18) | −0.00085 (12) | −0.00046 (12) | 0.00252 (12) |
| O1S | 0.0177 (5) | 0.0214 (5) | 0.0114 (5) | −0.0058 (4) | −0.0017 (4) | 0.0028 (4) |
| N1 | 0.0145 (6) | 0.0133 (6) | 0.0104 (5) | −0.0007 (4) | 0.0000 (4) | 0.0029 (4) |
| C2 | 0.0141 (6) | 0.0120 (6) | 0.0110 (6) | 0.0010 (5) | 0.0018 (5) | 0.0022 (5) |
| N2 | 0.0159 (6) | 0.0205 (6) | 0.0116 (6) | −0.0052 (5) | −0.0023 (5) | 0.0064 (5) |
| C3 | 0.0121 (6) | 0.0137 (6) | 0.0107 (6) | 0.0005 (5) | 0.0001 (5) | 0.0033 (5) |
| N3 | 0.0139 (6) | 0.0153 (6) | 0.0113 (6) | −0.0009 (4) | 0.0003 (4) | 0.0036 (4) |
| O3A | 0.0174 (5) | 0.0272 (6) | 0.0200 (5) | −0.0096 (4) | −0.0054 (4) | 0.0103 (5) |
| O3B | 0.0188 (5) | 0.0276 (6) | 0.0127 (5) | −0.0031 (4) | −0.0007 (4) | 0.0091 (4) |
| N4 | 0.0129 (5) | 0.0118 (5) | 0.0125 (6) | 0.0018 (4) | 0.0018 (4) | 0.0026 (4) |
| C5 | 0.0117 (6) | 0.0120 (6) | 0.0135 (6) | −0.0009 (5) | 0.0016 (5) | 0.0030 (5) |
| N5 | 0.0129 (5) | 0.0140 (6) | 0.0152 (6) | −0.0004 (4) | 0.0012 (4) | 0.0030 (5) |
| O5A | 0.0148 (5) | 0.0222 (5) | 0.0190 (5) | −0.0040 (4) | −0.0048 (4) | 0.0047 (4) |
| O5B | 0.0199 (5) | 0.0259 (6) | 0.0178 (5) | −0.0070 (4) | −0.0002 (4) | 0.0107 (4) |
| C6 | 0.0131 (6) | 0.0120 (6) | 0.0112 (6) | 0.0016 (5) | 0.0017 (5) | 0.0018 (5) |
| N6 | 0.0150 (6) | 0.0193 (6) | 0.0126 (6) | −0.0031 (5) | −0.0006 (5) | 0.0056 (5) |
| C1S | 0.0188 (7) | 0.0257 (8) | 0.0158 (7) | 0.0037 (6) | 0.0021 (6) | −0.0011 (6) |
| C2S | 0.0218 (8) | 0.0209 (8) | 0.0216 (8) | −0.0091 (6) | −0.0051 (6) | 0.0090 (6) |
Geometric parameters (Å, º) top
| S1S—O1S | 1.5174 (11) | N3—O3B | 1.2226 (16) |
| S1S—C2S | 1.7852 (16) | N3—O3A | 1.2347 (16) |
| S1S—C1S | 1.7855 (16) | N4—C5 | 1.3069 (19) |
| N1—C6 | 1.3438 (18) | C5—C6 | 1.4426 (19) |
| N1—C2 | 1.3458 (18) | C5—N5 | 1.4535 (17) |
| C2—N2 | 1.3251 (18) | N5—O5B | 1.2273 (17) |
| C2—C3 | 1.4405 (19) | N5—O5A | 1.2396 (16) |
| C3—N4 | 1.3018 (18) | C6—N6 | 1.3274 (19) |
| C3—N3 | 1.4555 (17) | | |
| | | |
| O1S—S1S—C2S | 105.12 (7) | O3A—N3—C3 | 117.79 (11) |
| O1S—S1S—C1S | 106.08 (7) | C3—N4—C5 | 119.07 (12) |
| C2S—S1S—C1S | 98.55 (9) | N4—C5—C6 | 122.24 (13) |
| C6—N1—C2 | 120.51 (12) | N4—C5—N5 | 114.15 (12) |
| N2—C2—N1 | 117.72 (13) | C6—C5—N5 | 123.61 (13) |
| N2—C2—C3 | 124.14 (13) | O5B—N5—O5A | 122.99 (12) |
| N1—C2—C3 | 118.14 (12) | O5B—N5—C5 | 118.81 (12) |
| N4—C3—C2 | 122.12 (13) | O5A—N5—C5 | 118.19 (12) |
| N4—C3—N3 | 114.65 (12) | N6—C6—N1 | 118.34 (13) |
| C2—C3—N3 | 123.22 (12) | N6—C6—C5 | 123.80 (13) |
| O3B—N3—O3A | 123.32 (12) | N1—C6—C5 | 117.87 (12) |
| O3B—N3—C3 | 118.88 (12) | | |
| | | |
| C6—N1—C2—N2 | 177.11 (13) | C3—N4—C5—C6 | −1.1 (2) |
| C6—N1—C2—C3 | −2.6 (2) | C3—N4—C5—N5 | 179.04 (12) |
| N2—C2—C3—N4 | −177.40 (13) | N4—C5—N5—O5B | −0.68 (19) |
| N1—C2—C3—N4 | 2.3 (2) | C6—C5—N5—O5B | 179.41 (13) |
| N2—C2—C3—N3 | 2.2 (2) | N4—C5—N5—O5A | −179.98 (12) |
| N1—C2—C3—N3 | −178.10 (12) | C6—C5—N5—O5A | 0.1 (2) |
| N4—C3—N3—O3B | 0.54 (19) | C2—N1—C6—N6 | −179.27 (12) |
| C2—C3—N3—O3B | −179.13 (13) | C2—N1—C6—C5 | 1.2 (2) |
| N4—C3—N3—O3A | −178.82 (13) | N4—C5—C6—N6 | −178.79 (13) |
| C2—C3—N3—O3A | 1.5 (2) | N5—C5—C6—N6 | 1.1 (2) |
| C2—C3—N4—C5 | −0.4 (2) | N4—C5—C6—N1 | 0.7 (2) |
| N3—C3—N4—C5 | 179.91 (12) | N5—C5—C6—N1 | −179.40 (12) |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| N2—H2A···O3A | 0.83 (2) | 2.10 (2) | 2.6703 (18) | 125.6 (17) |
| N2—H2A···O1Si | 0.83 (2) | 2.10 (2) | 2.7884 (17) | 139.9 (18) |
| N2—H2B···N1i | 0.80 (2) | 2.25 (2) | 3.0407 (18) | 170 (2) |
| N6—H6A···O5A | 0.86 (2) | 2.06 (2) | 2.6831 (18) | 128.8 (18) |
| N6—H6B···O1S | 0.82 (2) | 2.05 (2) | 2.8541 (17) | 171 (2) |
| Symmetry code: (i) −x+1, −y+1, −z+2. |
Experimental details
| Crystal data |
| Chemical formula | C4H4N6O4·C2H6OS |
| Mr | 278.26 |
| Crystal system, space group | Triclinic, P1 |
| Temperature (K) | 93 |
| a, b, c (Å) | 5.7817 (5), 8.1353 (8), 12.0270 (11) |
| α, β, γ (°) | 99.253 (2), 94.113 (2), 92.482 (2) |
| V (Å3) | 556.04 (9) |
| Z | 2 |
| Radiation type | Mo Kα |
| µ (mm−1) | 0.32 |
| Crystal size (mm) | 0.65 × 0.22 × 0.07 |
| |
| Data collection |
| Diffractometer | CCD area detector
diffractometer |
| Absorption correction | Integration
(Wuensch & Prewitt, 1965) |
| Tmin, Tmax | 0.919, 0.969 |
No. of measured, independent and
observed [I > 2σ(I)] reflections | 4175, 2638, 2505 |
| Rint | 0.031 |
| (sin θ/λ)max (Å−1) | 0.667 |
| |
| Refinement |
| R[F2 > 2σ(F2)], wR(F2), S | 0.035, 0.091, 1.14 |
| No. of reflections | 2638 |
| No. of parameters | 204 |
| H-atom treatment | H atoms treated by a mixture of independent and constrained refinement |
| Δρmax, Δρmin (e Å−3) | 0.44, −0.25 |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| N2—H2A···O3A | 0.83 (2) | 2.10 (2) | 2.6703 (18) | 125.6 (17) |
| N2—H2A···O1Si | 0.83 (2) | 2.10 (2) | 2.7884 (17) | 139.9 (18) |
| N2—H2B···N1i | 0.80 (2) | 2.25 (2) | 3.0407 (18) | 170 (2) |
| N6—H6A···O5A | 0.86 (2) | 2.06 (2) | 2.6831 (18) | 128.8 (18) |
| N6—H6B···O1S | 0.82 (2) | 2.05 (2) | 2.8541 (17) | 171 (2) |
| Symmetry code: (i) −x+1, −y+1, −z+2. |

 Subscribe to Acta Crystallographica Section E: Crystallographic Communications
Subscribe to Acta Crystallographica Section E: Crystallographic Communications
The full text of this article is available to subscribers to the journal.
If you have already registered and are using a computer listed in your registration details, please email
support@iucr.org for assistance.
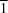 . It is a dimethyl sulfoxide (DMSO) solvate of the insensitive energetic compound 2,6-diamino-3,5-dinitro-1,4-pyrazine (ANPZ). The structure has been determined to obtain more accurate metrical parameters for the 2,6-diamino-3,5-dinitro-1,4-pyrazine moiety than those obtained from unsolvated crystals which are invariably twinned. The packing motif consists of two formula units linked by strong complementary hydrogen bonds between the ANPZ units with the two DMSO solvate molecules each linked by two bifurcated hydogen bonds to the two ANPZ units. This central motif is further linked into planar sheets by weaker interactions between the DMSO methyl H atoms and the O atoms from the nitro groups. This looser packing arrangement compared to ANPZ is reflected in a lower density (1.662 Mg m-3 versus 1.812 Mg m-3 for ANPZ).
. It is a dimethyl sulfoxide (DMSO) solvate of the insensitive energetic compound 2,6-diamino-3,5-dinitro-1,4-pyrazine (ANPZ). The structure has been determined to obtain more accurate metrical parameters for the 2,6-diamino-3,5-dinitro-1,4-pyrazine moiety than those obtained from unsolvated crystals which are invariably twinned. The packing motif consists of two formula units linked by strong complementary hydrogen bonds between the ANPZ units with the two DMSO solvate molecules each linked by two bifurcated hydogen bonds to the two ANPZ units. This central motif is further linked into planar sheets by weaker interactions between the DMSO methyl H atoms and the O atoms from the nitro groups. This looser packing arrangement compared to ANPZ is reflected in a lower density (1.662 Mg m-3 versus 1.812 Mg m-3 for ANPZ).
 journal menu
journal menu











![[Figure 1]](http://journals.iucr.org/e/issues/2001/08/00/wn6030/wn6030fig1thm.gif)
![[Figure 2]](http://journals.iucr.org/e/issues/2001/08/00/wn6030/wn6030fig2thm.gif)




The title compound, 2,6-diamino-3,5-dinitro-1,4-pyrazine dimethyl sulfoxide solvate, (I), is a dimethyl sulfoxide (DMSO) solvate of the insensitive energetic molecule 2,6-diamino-3,5-dinitro-1,4-pyrazine (ANPZ). ANPZ is a member of an important family of insensitive energetic compounds (TATB: Cady & Larson, 1965; Kolb & Rizzo, 1979; ANPZ: Gilardi & Butcher, 2001; ANPZO: Gilardi & Butcher, 2001), in which the molecules exhibit both multiple intramolecular and intermolecular hydrogen bonding interactions (TATB is 1,3,5-triamino-2,4,6-trinitrobenzene and ANPZO is 2,6-diamino-3,5-dinitropyrazine-1-oxide).
The intermolecular hydrogen-bonding interactions result in a planar sheet-like packing arrangement where the spacing between the layers resembles that of graphite. The significant feature of this family of energetic molecules is their insensitivity. Sensitivity is often tested via the drop height method, i.e. the height of the drop of a steel ball required to detonate the compound, with large values reflecting insensitivity. In such testing, in common with TATB, the benchmark compound as regards insensitivity, ANPZ has values which are so large they cannot be accurately measured, while ANPZO has a value of 117 cm (Pagoria et al., 1998). Thus ANPZ is much safer than other commonly used energetic compounds such as trinitrotoluene (80 cm) and HMX (32 cm). The primary motivating factor in determining the structure of the ANPZ solvate was to obtain more accurate metrical parameters for ANPZ as its crystals are invariably twinned (Gilardi & Butcher, 2001). The packing motif for the DMSO solvate consists of two ANPZ molecules linked by strong complementary hydrogen bonds and DMSO solvate molecules, each linked by two bifurcated hydrogen bonds to the ANPZ dimer. This central motif is further linked into planar sheets by weaker interactions between the DMSO methyl H atoms and the O atoms from the nitro groups. This packing arrangement is looser than that of ANPZ, and results in a lower density (1.662 Mg m-3 versus 1.812 Mg m-3 for ANPZ). Fig. 1 shows the structure and labeling scheme for the title compound, while Fig. 2 shows the packing arrangement. Hydrogen-bonding metrical parameters are given in Table 1.