Buy article online - an online subscription or single-article purchase is required to access this article.

Full three-dimensional diffuse scattering data have been recorded for both polymorphic forms [(I) and (II)] of aspirin and these data have been analysed using Monte Carlo computer modelling. The observed scattering in form (I) is well reproduced by a simple harmonic model of thermally induced displacements. The data for form (II) show, in addition to thermal diffuse scattering (TDS) similar to that in form (I), diffuse streaks originating from stacking fault-like defects as well as other effects that can be attributed to strain induced by these defects. The present study has provided strong evidence that the aspirin form (II) structure is a true polymorph with a structure quite distinct from that of form (I). The diffuse scattering evidence presented shows that crystals of form (II) are essentially composed of large single domains of the form (II) lattice with a relatively small volume fraction of intrinsic planar defects or faults comprising misoriented bilayers of molecular dimers. There is evidence of some local aggregation of these defect bilayers to form small included regions of the form (I) structure. Evidence is also presented that shows that the strain effects arise from the mismatch of molecular packing between the defect region and the surrounding form (II) lattice. This occurs at the edges of the planar defects in the
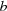
direction only.
Supporting information
CCDC references: 810889; 1101022
Cell refinement: HKL SCALEPACK (Otwinowski & Minor 1997); data reduction: HKL DENZO and SCALEPACK (Otwinowski & Minor 1997); program(s) used to solve structure: known; program(s) used to refine structure: SHELXL97 (Sheldrick, 1997); molecular graphics: Xtal3.7 (Hall et al., 2001); software used to prepare material for publication: WinGX publication routines (Farrugia, 1999).
Crystal data top
| C9H8O4 | F(000) = 376 |
| Mr = 180.15 | Dx = 1.394 Mg m−3 |
| Monoclinic, P21/c | Mo Kα radiation, λ = 0.71073 Å |
| Hall symbol: -P 2ybc | Cell parameters from 8462 reflections |
| a = 12.2696 (5) Å | θ = 27.5–2.6° |
| b = 6.5575 (3) Å | µ = 0.11 mm−1 |
| c = 11.4960 (4) Å | T = 300 K |
| β = 68.163 (2)° | Prism, colourless |
| V = 858.58 (6) Å3 | 0.4 × 0.2 × 0.1 mm |
| Z = 4 | |
Data collection top
KappaCCD
diffractometer | 1853 independent reflections |
| Radiation source: fine-focus sealed tube | 1425 reflections with I > 2σ(I) |
| Graphite monochromator | Rint = 0.031 |
| Detector resolution: 9 pixels mm-1 | θmax = 27.2°, θmin = 3.6° |
| CCD scans | h = −15→15 |
Absorption correction: integration
Gaussian integration (Coppens, 1970) | k = −5→8 |
| Tmin = 0.968, Tmax = 0.986 | l = −14→14 |
| 6924 measured reflections | |
Refinement top
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.043 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.129 | H-atom parameters constrained |
| S = 1.09 | w = 1/[σ2(Fo2) + (0.065P)2 + 0.127P]
where P = (Fo2 + 2Fc2)/3 |
| 1853 reflections | (Δ/σ)max < 0.001 |
| 119 parameters | Δρmax = 0.23 e Å−3 |
| 0 restraints | Δρmin = −0.20 e Å−3 |
Special details top
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes)
are estimated using the full covariance matrix. The cell e.s.d.'s are taken
into account individually in the estimation of e.s.d.'s in distances, angles
and torsion angles; correlations between e.s.d.'s in cell parameters are only
used when they are defined by crystal symmetry. An approximate (isotropic)
treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s.
planes. |
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor
wR and goodness of fit S are based on F2, conventional
R-factors R are based on F, with F set to zero for
negative F2. The threshold expression of F2 >
σ(F2) is used only for calculating R-factors(gt) etc.
and is not relevant to the choice of reflections for refinement.
R-factors based on F2 are statistically about twice as large
as those based on F, and R- factors based on ALL data will be
even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| C1 | 0.15361 (12) | 0.5634 (2) | −0.00763 (12) | 0.0422 (4) | |
| C2 | 0.24609 (13) | 0.4867 (2) | −0.11120 (12) | 0.0445 (4) | |
| C3 | 0.29856 (14) | 0.3035 (3) | −0.10485 (15) | 0.0541 (4) | |
| H14 | 0.3591 | 0.2535 | −0.1751 | 0.065* | |
| C4 | 0.26148 (15) | 0.1939 (3) | 0.00552 (16) | 0.0584 (4) | |
| H15 | 0.2976 | 0.0709 | 0.0095 | 0.070* | |
| C5 | 0.17144 (16) | 0.2657 (3) | 0.10958 (16) | 0.0552 (4) | |
| H16 | 0.1468 | 0.1920 | 0.1840 | 0.066* | |
| C6 | 0.11776 (14) | 0.4480 (2) | 0.10276 (13) | 0.0487 (4) | |
| H17 | 0.0564 | 0.4953 | 0.1732 | 0.058* | |
| C7 | 0.09012 (13) | 0.7565 (2) | −0.00660 (13) | 0.0432 (4) | |
| O8 | 0.00975 (10) | 0.81103 (19) | 0.09154 (10) | 0.0604 (4) | |
| O9 | 0.12013 (10) | 0.85938 (19) | −0.10910 (10) | 0.0580 (3) | |
| H18 | 0.0796 | 0.9624 | −0.0975 | 0.087* | |
| O10 | 0.28510 (9) | 0.58717 (18) | −0.22724 (8) | 0.0494 (3) | |
| C11 | 0.36529 (13) | 0.7386 (3) | −0.24386 (13) | 0.0484 (4) | |
| O12 | 0.40430 (11) | 0.7814 (2) | −0.16633 (10) | 0.0627 (4) | |
| C13 | 0.39556 (17) | 0.8367 (4) | −0.36860 (16) | 0.0710 (6) | |
| H19 | 0.4527 | 0.9420 | −0.3783 | 0.107* | |
| H20 | 0.3262 | 0.8952 | −0.3747 | 0.107* | |
| H21 | 0.4274 | 0.7364 | −0.4332 | 0.107* | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| C1 | 0.0385 (7) | 0.0489 (8) | 0.0387 (7) | −0.0031 (6) | −0.0139 (6) | −0.0013 (6) |
| C2 | 0.0417 (8) | 0.0524 (9) | 0.0382 (7) | −0.0042 (6) | −0.0135 (6) | −0.0022 (6) |
| C3 | 0.0480 (9) | 0.0567 (10) | 0.0522 (9) | 0.0053 (7) | −0.0125 (7) | −0.0090 (7) |
| C4 | 0.0576 (10) | 0.0511 (10) | 0.0664 (10) | 0.0056 (8) | −0.0229 (8) | 0.0009 (8) |
| C5 | 0.0554 (10) | 0.0561 (10) | 0.0522 (9) | −0.0040 (8) | −0.0176 (7) | 0.0105 (7) |
| C6 | 0.0456 (8) | 0.0551 (10) | 0.0420 (8) | −0.0014 (7) | −0.0124 (6) | 0.0026 (7) |
| C7 | 0.0400 (8) | 0.0524 (9) | 0.0354 (7) | −0.0028 (6) | −0.0117 (6) | −0.0005 (6) |
| O8 | 0.0624 (7) | 0.0663 (8) | 0.0415 (6) | 0.0164 (6) | −0.0067 (5) | 0.0004 (5) |
| O9 | 0.0574 (7) | 0.0617 (7) | 0.0461 (6) | 0.0101 (6) | −0.0091 (5) | 0.0100 (5) |
| O10 | 0.0477 (6) | 0.0635 (7) | 0.0343 (5) | −0.0028 (5) | −0.0122 (4) | −0.0018 (4) |
| C11 | 0.0409 (8) | 0.0624 (10) | 0.0370 (7) | 0.0025 (7) | −0.0088 (6) | 0.0003 (7) |
| O12 | 0.0619 (8) | 0.0819 (9) | 0.0463 (6) | −0.0176 (6) | −0.0226 (6) | 0.0077 (6) |
| C13 | 0.0651 (11) | 0.0989 (15) | 0.0470 (9) | −0.0108 (11) | −0.0186 (8) | 0.0211 (9) |
Geometric parameters (Å, º) top
| C1—C2 | 1.3974 (19) | C6—H17 | 0.9300 |
| C1—C6 | 1.400 (2) | C7—O8 | 1.2432 (17) |
| C1—C7 | 1.485 (2) | C7—O9 | 1.2869 (17) |
| C2—C3 | 1.378 (2) | O9—H18 | 0.8200 |
| C2—O10 | 1.4027 (17) | O10—C11 | 1.361 (2) |
| C3—C4 | 1.380 (2) | C11—O12 | 1.1915 (19) |
| C3—H14 | 0.9300 | C11—C13 | 1.487 (2) |
| C4—C5 | 1.375 (2) | C13—H19 | 0.9600 |
| C4—H15 | 0.9300 | C13—H20 | 0.9600 |
| C5—C6 | 1.381 (2) | C13—H21 | 0.9600 |
| C5—H16 | 0.9300 | | |
| | | |
| C2—C1—C6 | 117.37 (14) | C5—C6—H17 | 119.2 |
| C2—C1—C7 | 124.81 (13) | C1—C6—H17 | 119.2 |
| C6—C1—C7 | 117.82 (12) | O8—C7—O9 | 122.72 (14) |
| C3—C2—C1 | 121.05 (14) | O8—C7—C1 | 119.28 (13) |
| C3—C2—O10 | 117.33 (13) | O9—C7—C1 | 117.99 (12) |
| C1—C2—O10 | 121.54 (14) | C7—O9—H18 | 109.5 |
| C2—C3—C4 | 120.13 (14) | C11—O10—C2 | 116.72 (11) |
| C2—C3—H14 | 119.9 | O12—C11—O10 | 122.52 (14) |
| C4—C3—H14 | 119.9 | O12—C11—C13 | 126.41 (16) |
| C5—C4—C3 | 120.37 (16) | O10—C11—C13 | 111.07 (14) |
| C5—C4—H15 | 119.8 | C11—C13—H19 | 109.5 |
| C3—C4—H15 | 119.8 | C11—C13—H20 | 109.5 |
| C4—C5—C6 | 119.51 (15) | H19—C13—H20 | 109.5 |
| C4—C5—H16 | 120.2 | C11—C13—H21 | 109.5 |
| C6—C5—H16 | 120.2 | H19—C13—H21 | 109.5 |
| C5—C6—C1 | 121.55 (14) | H20—C13—H21 | 109.5 |

 Subscribe to Acta Crystallographica Section B: Structural Science, Crystal Engineering and Materials
Subscribe to Acta Crystallographica Section B: Structural Science, Crystal Engineering and Materials
The full text of this article is available to subscribers to the journal.
If you have already registered and are using a computer listed in your registration details, please email
support@iucr.org for assistance.
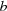 direction only.
direction only.
 journal menu
journal menu









 Subscribe to Acta Crystallographica Section B: Structural Science, Crystal Engineering and Materials
Subscribe to Acta Crystallographica Section B: Structural Science, Crystal Engineering and Materials


