Buy article online - an online subscription or single-article purchase is required to access this article.
Although molecules of the title compound, 3,4-dichloro-2,5-dihydrofuran-2,5-dione (dichloromaleic anhydride, C
4Cl
2O
3), (I), possess approximate non-crystallographic
C2v symmetry, the two chlorine substituents deviate from the ring plane. Their deviations are in the same direction, but with values of 0.0356 (17) and 0.0167 (17) Å, they differ significantly in magnitude. The closest intermolecular contact is of 2.888 (2) Å between a carbonyl O atom and the C atom of a carbonyl group, with the O

C direction orthogonal to the C=O bond [O5

C2
i=O2
i 93.6 (2)°; symmetry code: (i)
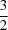
-
x, -½ +
y, z]. These contacts form infinite chains of molecules running parallel to the crystallographic
b direction.
Supporting information
CCDC reference: 146107
Data collection: SMART (Bruker, 1998); cell refinement: SMART; data reduction: SAINT (Bruker, 1999); program(s) used to solve structure: SHELXTL (Bruker, 1997); program(s) used to refine structure: SHELXL97 (Sheldrick, 1997); software used to prepare material for publication: SHELXL97 and PLATON (Spek, 1999).
Crystal data top
| C4Cl2O3 | F(000) = 656 |
| Mr = 166.94 | Dx = 1.941 Mg m−3 |
| Orthorhombic, Pbca | Mo Kα radiation, λ = 0.71073 Å |
| Hall symbol: -P 2ac 2ab | Cell parameters from 5130 reflections |
| a = 10.6397 (6) Å | θ = 3.1–28.6° |
| b = 8.2558 (5) Å | µ = 1.05 mm−1 |
| c = 13.0057 (7) Å | T = 150 K |
| V = 1142.41 (11) Å3 | Block, colourless |
| Z = 8 | 0.32 × 0.32 × 0.25 mm |
Data collection top
Bruker SMART CCD area-detector
diffractometer | 1372 independent reflections |
| Radiation source: fine-focus sealed tube | 1261 reflections with I > 2σ(I) |
| Graphite monochromator | Rint = 0.013 |
| ω scans | θmax = 28.6°, θmin = 3.1° |
Absorption correction: multi-scan
(SADABS; Bruker, 1996) | h = 0→14 |
| Tmin = 0.730, Tmax = 0.779 | k = 0→11 |
| 7980 measured reflections | l = 0→17 |
Refinement top
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | w = 1/[σ2(Fo2) + (0.041P)2 + 0.34P]
where P = (Fo2 + 2Fc2)/3 |
| R[F2 > 2σ(F2)] = 0.023 | (Δ/σ)max = 0.001 |
| wR(F2) = 0.067 | Δρmax = 0.35 e Å−3 |
| S = 1.04 | Δρmin = −0.21 e Å−3 |
| 1372 reflections | Extinction correction: SHELXL97, Fc*=kFc[1+0.001xFc2λ3/sin(2θ)]-1/4 |
| 83 parameters | Extinction coefficient: 0.0056 (7) |
| 0 restraints | |
Special details top
Geometry. Least-squares planes (x,y,z in crystal coordinates) and
deviations from them (* indicates atom used to define plane) 5.8224 (0.0054) x + 6.8975 (0.0028) y - 0.6529 (0.0066) z
= 4.1943 (0.0036) * -0.0007 (0.0006) O1 * 0.0023 (0.0006) C2 * -0.0031 (0.0007) C3 * 0.0028
(0.0007) C4 * -0.0012 (0.0006) C5 - 0.0039 (0.0017) O2 - 0.0023 (0.0018) O5 -
0.0356 (0.0017) Cl3 - 0.0167 (0.0017) Cl4 Rms deviation of fitted atoms = 0.0022
___________________________________________________________________ Short Non-Hydrogen Inter-Molecular Contacts Contact-Nr Atom I [ARU] Atom J [ARU] d(I—J) Del
——————————————————————- 1 O(2)
[1555.01] ··· C(4) [4655.01] = 3.019 - 0.20 2 O(2) [1555.01] ··· C(5)
[4655.01] = 3.011 - 0.21 3 O(5) [1555.01] ··· C(2) [7645.01] = 2.888 - 0.33 [4655.01] = 1 - x,1/2 + y,1/2 - z [7645.01] = 3/2 -
x,-1/2 + y,z |
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor
wR and goodness of fit S are based on F2, conventional
R-factors R are based on F, with F set to zero for
negative F2. The threshold expression of F2 >
σ(F2) is used only for calculating R-factors(gt) etc.
and is not relevant to the choice of reflections for refinement.
R-factors based on F2 are statistically about twice as large
as those based on F, and R- factors based on ALL data will be
even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| O1 | 0.62393 (8) | 0.09767 (10) | 0.17283 (6) | 0.0261 (2) | |
| C2 | 0.53423 (10) | 0.17933 (13) | 0.23110 (9) | 0.0241 (2) | |
| O2 | 0.44778 (10) | 0.24782 (10) | 0.19319 (8) | 0.0319 (2) | |
| C3 | 0.56901 (11) | 0.15962 (13) | 0.34126 (9) | 0.0232 (2) | |
| Cl3 | 0.48172 (3) | 0.23767 (4) | 0.43703 (3) | 0.03491 (12) | |
| C4 | 0.67404 (10) | 0.07227 (13) | 0.34605 (8) | 0.0223 (2) | |
| Cl4 | 0.75628 (3) | 0.00979 (4) | 0.44921 (2) | 0.03129 (12) | |
| C5 | 0.71187 (11) | 0.02966 (14) | 0.23941 (9) | 0.0235 (2) | |
| O5 | 0.79899 (9) | −0.04691 (11) | 0.20904 (7) | 0.0310 (2) | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| O1 | 0.0300 (4) | 0.0266 (4) | 0.0217 (4) | 0.0028 (3) | −0.0014 (3) | −0.0012 (3) |
| C2 | 0.0239 (5) | 0.0197 (5) | 0.0287 (6) | −0.0025 (4) | 0.0006 (4) | −0.0003 (4) |
| O2 | 0.0264 (5) | 0.0297 (5) | 0.0397 (6) | 0.0017 (3) | −0.0057 (4) | 0.0047 (4) |
| C3 | 0.0244 (5) | 0.0210 (5) | 0.0241 (5) | −0.0036 (4) | 0.0034 (4) | −0.0017 (4) |
| Cl3 | 0.0336 (2) | 0.0363 (2) | 0.0348 (2) | −0.00032 (12) | 0.01382 (12) | −0.00629 (12) |
| C4 | 0.0235 (5) | 0.0226 (5) | 0.0209 (5) | −0.0039 (4) | −0.0003 (4) | 0.0011 (4) |
| Cl4 | 0.03173 (19) | 0.0370 (2) | 0.02518 (19) | −0.00352 (12) | −0.00609 (11) | 0.00644 (11) |
| C5 | 0.0250 (5) | 0.0203 (5) | 0.0253 (6) | −0.0020 (4) | −0.0001 (4) | −0.0001 (4) |
| O5 | 0.0303 (5) | 0.0299 (4) | 0.0327 (5) | 0.0060 (4) | 0.0050 (4) | −0.0028 (4) |
Geometric parameters (Å, º) top
| O1—C2 | 1.3927 (14) | C3—Cl3 | 1.6820 (12) |
| O1—C5 | 1.3929 (14) | C4—C5 | 1.4864 (16) |
| C2—O2 | 1.1870 (15) | C4—Cl4 | 1.6829 (11) |
| C2—C3 | 1.4886 (16) | C5—O5 | 1.1895 (15) |
| C3—C4 | 1.3314 (17) | | |
| | | |
| C2—O1—C5 | 108.49 (9) | C3—C4—C5 | 108.17 (10) |
| O2—C2—O1 | 122.39 (11) | C3—C4—Cl4 | 129.77 (9) |
| O2—C2—C3 | 130.13 (11) | C5—C4—Cl4 | 122.05 (9) |
| O1—C2—C3 | 107.48 (9) | O5—C5—O1 | 122.09 (11) |
| C4—C3—C2 | 108.23 (10) | O5—C5—C4 | 130.29 (11) |
| C4—C3—Cl3 | 129.51 (9) | O1—C5—C4 | 107.62 (10) |
| C2—C3—Cl3 | 122.25 (9) | | |
| | | |
| C5—O1—C2—O2 | −179.59 (11) | C2—C3—C4—Cl4 | −179.23 (9) |
| C5—O1—C2—C3 | −0.27 (11) | Cl3—C3—C4—Cl4 | −0.15 (18) |
| O2—C2—C3—C4 | 179.77 (12) | C2—O1—C5—O5 | −179.93 (11) |
| O1—C2—C3—C4 | 0.53 (13) | C2—O1—C5—C4 | −0.05 (11) |
| O2—C2—C3—Cl3 | 0.61 (18) | C3—C4—C5—O5 | −179.74 (13) |
| O1—C2—C3—Cl3 | −178.63 (7) | Cl4—C4—C5—O5 | −0.94 (18) |
| C2—C3—C4—C5 | −0.55 (13) | C3—C4—C5—O1 | 0.39 (12) |
| Cl3—C3—C4—C5 | 178.53 (9) | Cl4—C4—C5—O1 | 179.19 (8) |

 Subscribe to Acta Crystallographica Section C: Structural Chemistry
Subscribe to Acta Crystallographica Section C: Structural Chemistry
The full text of this article is available to subscribers to the journal.
If you have already registered and are using a computer listed in your registration details, please email
support@iucr.org for assistance.
 C direction orthogonal to the C=O bond [O5
C direction orthogonal to the C=O bond [O5 C2i=O2i 93.6 (2)°; symmetry code: (i)
C2i=O2i 93.6 (2)°; symmetry code: (i) 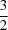 - x, -½ + y, z]. These contacts form infinite chains of molecules running parallel to the crystallographic b direction.
- x, -½ + y, z]. These contacts form infinite chains of molecules running parallel to the crystallographic b direction.  C direction orthogonal to the C=O bond [O5
C direction orthogonal to the C=O bond [O5 C2i=O2i 93.6 (2)°; symmetry code: (i)
C2i=O2i 93.6 (2)°; symmetry code: (i) 
 journal menu
journal menu









 Subscribe to Acta Crystallographica Section C: Structural Chemistry
Subscribe to Acta Crystallographica Section C: Structural Chemistry


