Buy article online - an online subscription or single-article purchase is required to access this article.
Isoguanine, an analogue of guanine, is of intrinsic interest as a noncanonical nucleobase. The crystal structure of isoguaninium chloride (systematic name: 6-amino-2-oxo-1
H,7
H-purin-3-ium chloride), C
5H
6N
5O
+·Cl
−, has been determined by single-crystal X-ray diffraction. Structure analysis was supported by electrostatic interaction energy (
Ees) calculations based on charge density reconstructed with the UBDB databank. In the structure, two kinds of molecular tapes are observed, one parallel to (010) and the other parallel to (50
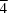
). The tapes are formed by dimers of isoguaninium cations interacting with chloride anions.
Ees analysis indicates that cations in one kind of tape are oriented so as to minimize repulsive electrostatic interactions.
Supporting information
CCDC reference: 1569459
Data collection: CrysAlis PRO (Agilent, 2014); cell refinement: CrysAlis PRO (Agilent, 2014); data reduction: CrysAlis PRO (Agilent, 2014); program(s) used to solve structure: SHELXS97 (Sheldrick, 2008); program(s) used to refine structure: SHELXL2017 (Sheldrick, 2015); molecular graphics: OLEX2 (Dolomanov et al., 2009); software used to prepare material for publication: OLEX2 (Dolomanov et al., 2009).
6-Amino-2-oxo-1
H,7
H-purin-3-ium chloride
top
Crystal data top
| C5H6N5O+·Cl− | F(000) = 768 |
| Mr = 187.60 | Dx = 1.685 Mg m−3 |
| Monoclinic, P21/c | Cu Kα radiation, λ = 1.54184 Å |
| a = 14.0359 (6) Å | Cell parameters from 11926 reflections |
| b = 13.6930 (5) Å | θ = 4.5–76.2° |
| c = 7.7026 (3) Å | µ = 4.25 mm−1 |
| β = 92.754 (4)° | T = 100 K |
| V = 1478.68 (10) Å3 | Needle, colourless |
| Z = 8 | 0.73 × 0.12 × 0.09 mm |
Data collection top
Agilent SuperNova Dual Source
diffractometer with an Atlas detector | 5522 independent reflections |
| Radiation source: sealed X-ray tube, SuperNova (Cu) X-ray Source | 4803 reflections with I > 2σ(I) |
| Mirror monochromator | Rint = 0.132 |
| Detector resolution: 5.2195 pixels mm-1 | θmax = 76.9°, θmin = 3.2° |
| ω scans | h = −17→17 |
Absorption correction: multi-scan
(CrysAlis PRO; Agilent, 2014) | k = −17→17 |
| Tmin = 0.465, Tmax = 1.000 | l = −9→9 |
| 42770 measured reflections | |
Refinement top
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Hydrogen site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.062 | All H-atom parameters refined |
| wR(F2) = 0.192 | w = 1/[σ2(Fo2) + (0.1497P)2 + 0.6578P]
where P = (Fo2 + 2Fc2)/3 |
| S = 1.06 | (Δ/σ)max < 0.001 |
| 5522 reflections | Δρmax = 0.72 e Å−3 |
| 266 parameters | Δρmin = −0.60 e Å−3 |
| 0 restraints | |
Special details top
Experimental. Single crystals of iGHCl were selected and mounted with paratone-N
oil on a MiTeGen micro-mount. The crystal was kept at 100.0 (1) K during
diffraction data collection on a Rigaku Oxford Diffraction (former Agilent
Technologies) SuperNova four-circle diffractometer with a Cu
Kα radiation and Atlas detector. The temperature was
controlled with an Oxford Cryosystems low-temperature nitrogen gas-flow device
(Cryostream Plus). The crystal was positioned 74 mm from the detector. A total
of 5060 frames were collected in 49 runs using ω scan with a rotation width
of 1.0°. The exposure time was in the range of 1–5 s. The determination of
unit-cell parameters, integration of reflection intensities and data
reduction, including a multiscan absorption correction, were performed
using CrysAlis PRO (Agilent, 2014). |
Geometry. All esds (except the esd in the dihedral angle between two l.s. planes)
are estimated using the full covariance matrix. The cell esds are taken
into account individually in the estimation of esds in distances, angles
and torsion angles; correlations between esds in cell parameters are only
used when they are defined by crystal symmetry. An approximate (isotropic)
treatment of cell esds is used for estimating esds involving l.s. planes. |
Refinement. Refined as a 2-component twin |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| Cl1 | 0.32116 (6) | 0.36689 (7) | 0.13649 (13) | 0.0208 (3) | |
| Cl2 | −0.20744 (6) | 0.17005 (6) | −0.53743 (13) | 0.0202 (3) | |
| O1 | 0.28818 (18) | 0.3810 (2) | −0.3447 (4) | 0.0224 (6) | |
| N1 | 0.4323 (2) | 0.3775 (2) | −0.1963 (4) | 0.0151 (6) | |
| N3 | 0.4224 (2) | 0.3789 (2) | −0.5007 (4) | 0.0167 (7) | |
| N7 | 0.6673 (2) | 0.3796 (2) | −0.3906 (5) | 0.0163 (7) | |
| N9 | 0.5746 (2) | 0.3807 (2) | −0.6363 (4) | 0.0174 (6) | |
| N10 | 0.5733 (2) | 0.3759 (2) | −0.0266 (5) | 0.0183 (7) | |
| C2 | 0.3741 (2) | 0.3795 (3) | −0.3490 (5) | 0.0172 (7) | |
| C4 | 0.5196 (2) | 0.3785 (3) | −0.4958 (5) | 0.0145 (7) | |
| C5 | 0.5741 (2) | 0.3775 (3) | −0.3414 (5) | 0.0162 (7) | |
| C6 | 0.5293 (2) | 0.3771 (3) | −0.1807 (5) | 0.0157 (8) | |
| C8 | 0.6635 (2) | 0.3806 (3) | −0.5653 (6) | 0.0170 (7) | |
| H1 | 0.403 (4) | 0.376 (4) | −0.096 (8) | 0.028 (13)* | |
| H3 | 0.388 (3) | 0.377 (3) | −0.610 (7) | 0.022 (12)* | |
| H7 | 0.715 (4) | 0.383 (4) | −0.330 (8) | 0.040 (16)* | |
| H8 | 0.721 (4) | 0.385 (4) | −0.634 (7) | 0.038 (15)* | |
| H10A | 0.637 (3) | 0.367 (3) | −0.024 (7) | 0.021 (12)* | |
| H10B | 0.539 (4) | 0.371 (4) | 0.064 (7) | 0.026 (13)* | |
| O2 | −0.12492 (17) | 0.42561 (19) | −0.4301 (4) | 0.0217 (6) | |
| N11 | 0.0093 (2) | 0.4414 (2) | −0.2529 (4) | 0.0159 (6) | |
| N13 | −0.0473 (2) | 0.2868 (2) | −0.3411 (4) | 0.0156 (6) | |
| N17 | 0.15561 (19) | 0.2389 (2) | −0.0712 (4) | 0.0149 (6) | |
| N19 | 0.0456 (2) | 0.1495 (2) | −0.2204 (4) | 0.0148 (6) | |
| N20 | 0.1439 (2) | 0.4653 (2) | −0.0703 (5) | 0.0196 (7) | |
| C12 | −0.0590 (2) | 0.3861 (3) | −0.3468 (5) | 0.0169 (7) | |
| C14 | 0.0260 (2) | 0.2465 (3) | −0.2420 (5) | 0.0142 (7) | |
| C15 | 0.0924 (2) | 0.3033 (3) | −0.1533 (5) | 0.0157 (7) | |
| C16 | 0.0853 (2) | 0.4063 (3) | −0.1548 (5) | 0.0156 (7) | |
| C18 | 0.1249 (2) | 0.1489 (3) | −0.1165 (5) | 0.0166 (7) | |
| H11 | 0.002 (3) | 0.506 (4) | −0.267 (6) | 0.026 (12)* | |
| H13 | −0.090 (3) | 0.252 (3) | −0.392 (6) | 0.013 (10)* | |
| H17 | 0.206 (3) | 0.253 (3) | −0.009 (6) | 0.012 (9)* | |
| H18 | 0.153 (3) | 0.088 (3) | −0.075 (6) | 0.016 (10)* | |
| H20A | 0.196 (3) | 0.443 (4) | −0.015 (6) | 0.024 (12)* | |
| H20B | 0.139 (4) | 0.531 (5) | −0.075 (8) | 0.053 (18)* | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| Cl1 | 0.0167 (4) | 0.0303 (5) | 0.0150 (5) | −0.0023 (3) | −0.0016 (3) | 0.0001 (4) |
| Cl2 | 0.0193 (4) | 0.0157 (4) | 0.0249 (5) | 0.0000 (3) | −0.0060 (3) | −0.0008 (3) |
| O1 | 0.0158 (12) | 0.0296 (15) | 0.0216 (17) | −0.0011 (10) | −0.0012 (11) | −0.0014 (12) |
| N1 | 0.0137 (13) | 0.0227 (16) | 0.0089 (16) | −0.0003 (11) | −0.0013 (12) | −0.0004 (12) |
| N3 | 0.0157 (14) | 0.0240 (16) | 0.0102 (17) | −0.0017 (11) | −0.0026 (12) | −0.0008 (12) |
| N7 | 0.0144 (14) | 0.0213 (16) | 0.0131 (17) | 0.0007 (11) | 0.0003 (12) | −0.0007 (13) |
| N9 | 0.0173 (13) | 0.0202 (15) | 0.0147 (17) | 0.0017 (11) | 0.0006 (12) | 0.0018 (13) |
| N10 | 0.0173 (14) | 0.0253 (17) | 0.0120 (17) | −0.0002 (11) | −0.0021 (12) | −0.0003 (13) |
| C2 | 0.0184 (16) | 0.0193 (17) | 0.014 (2) | −0.0003 (13) | 0.0030 (14) | −0.0012 (15) |
| C4 | 0.0162 (15) | 0.0139 (16) | 0.0133 (19) | −0.0011 (12) | −0.0008 (13) | 0.0003 (13) |
| C5 | 0.0140 (15) | 0.0177 (17) | 0.017 (2) | 0.0005 (12) | −0.0008 (14) | −0.0004 (14) |
| C6 | 0.0157 (15) | 0.0126 (16) | 0.019 (2) | −0.0002 (12) | −0.0004 (14) | 0.0016 (14) |
| C8 | 0.0168 (15) | 0.0173 (17) | 0.017 (2) | 0.0004 (12) | 0.0026 (14) | −0.0006 (14) |
| O2 | 0.0193 (11) | 0.0153 (12) | 0.0295 (17) | 0.0016 (9) | −0.0085 (11) | 0.0013 (11) |
| N11 | 0.0186 (13) | 0.0103 (14) | 0.0185 (17) | 0.0000 (10) | −0.0022 (12) | 0.0008 (11) |
| N13 | 0.0154 (12) | 0.0131 (14) | 0.0177 (17) | −0.0010 (11) | −0.0042 (11) | −0.0012 (12) |
| N17 | 0.0129 (12) | 0.0156 (15) | 0.0159 (17) | −0.0004 (10) | −0.0027 (11) | −0.0012 (12) |
| N19 | 0.0156 (13) | 0.0132 (14) | 0.0155 (17) | 0.0008 (10) | −0.0005 (11) | −0.0014 (12) |
| N20 | 0.0189 (14) | 0.0154 (15) | 0.0239 (19) | −0.0003 (11) | −0.0054 (13) | −0.0024 (13) |
| C12 | 0.0171 (15) | 0.0158 (17) | 0.017 (2) | 0.0002 (12) | −0.0016 (13) | 0.0013 (14) |
| C14 | 0.0153 (14) | 0.0159 (16) | 0.0113 (18) | 0.0016 (13) | 0.0008 (12) | −0.0001 (13) |
| C15 | 0.0153 (14) | 0.0142 (17) | 0.018 (2) | 0.0002 (12) | 0.0010 (13) | 0.0002 (14) |
| C16 | 0.0146 (14) | 0.0135 (16) | 0.019 (2) | 0.0003 (12) | 0.0027 (13) | 0.0006 (14) |
| C18 | 0.0181 (15) | 0.0131 (16) | 0.019 (2) | 0.0020 (12) | 0.0055 (14) | −0.0012 (14) |
Geometric parameters (Å, º) top
| O1—C2 | 1.208 (4) | O2—C12 | 1.226 (4) |
| N1—C2 | 1.399 (5) | N11—C12 | 1.397 (5) |
| N1—C6 | 1.361 (4) | N11—C16 | 1.365 (4) |
| N1—H1 | 0.89 (6) | N11—H11 | 0.90 (6) |
| N3—C2 | 1.378 (5) | N13—C12 | 1.369 (5) |
| N3—C4 | 1.363 (4) | N13—C14 | 1.368 (4) |
| N3—H3 | 0.95 (5) | N13—H13 | 0.85 (4) |
| N7—C5 | 1.379 (4) | N17—C15 | 1.383 (4) |
| N7—C8 | 1.344 (5) | N17—C18 | 1.346 (5) |
| N7—H7 | 0.80 (6) | N17—H17 | 0.86 (4) |
| N9—C4 | 1.359 (5) | N19—C14 | 1.365 (5) |
| N9—C8 | 1.338 (4) | N19—C18 | 1.339 (5) |
| N10—C6 | 1.312 (5) | N20—C16 | 1.304 (5) |
| N10—H10A | 0.91 (5) | N20—H20A | 0.88 (5) |
| N10—H10B | 0.87 (5) | N20—H20B | 0.91 (7) |
| C4—C5 | 1.383 (5) | C14—C15 | 1.370 (5) |
| C5—C6 | 1.415 (5) | C15—C16 | 1.413 (5) |
| C8—H8 | 0.98 (6) | C18—H18 | 0.97 (5) |
| | | |
| C2—N1—H1 | 117 (3) | C12—N11—H11 | 114 (3) |
| C6—N1—C2 | 127.9 (3) | C16—N11—C12 | 126.5 (3) |
| C6—N1—H1 | 115 (3) | C16—N11—H11 | 120 (3) |
| C2—N3—H3 | 120 (3) | C12—N13—H13 | 117 (3) |
| C4—N3—C2 | 120.6 (3) | C14—N13—C12 | 120.3 (3) |
| C4—N3—H3 | 119 (3) | C14—N13—H13 | 122 (3) |
| C5—N7—H7 | 128 (4) | C15—N17—H17 | 127 (3) |
| C8—N7—C5 | 106.4 (3) | C18—N17—C15 | 106.0 (3) |
| C8—N7—H7 | 125 (4) | C18—N17—H17 | 126 (3) |
| C8—N9—C4 | 103.2 (3) | C18—N19—C14 | 103.7 (3) |
| C6—N10—H10A | 116 (3) | C16—N20—H20A | 120 (3) |
| C6—N10—H10B | 118 (3) | C16—N20—H20B | 123 (4) |
| H10A—N10—H10B | 124 (4) | H20A—N20—H20B | 116 (5) |
| O1—C2—N1 | 121.3 (4) | O2—C12—N11 | 120.9 (3) |
| O1—C2—N3 | 123.7 (4) | O2—C12—N13 | 122.8 (3) |
| N3—C2—N1 | 114.9 (3) | N13—C12—N11 | 116.3 (3) |
| N3—C4—C5 | 122.3 (3) | N13—C14—C15 | 121.6 (3) |
| N9—C4—N3 | 125.7 (3) | N19—C14—N13 | 127.1 (3) |
| N9—C4—C5 | 111.9 (3) | N19—C14—C15 | 111.3 (3) |
| N7—C5—C4 | 104.8 (3) | N17—C15—C16 | 133.1 (3) |
| N7—C5—C6 | 135.1 (4) | C14—C15—N17 | 105.8 (3) |
| C4—C5—C6 | 120.1 (3) | C14—C15—C16 | 121.1 (3) |
| N1—C6—C5 | 114.1 (3) | N11—C16—C15 | 114.1 (3) |
| N10—C6—N1 | 120.4 (4) | N20—C16—N11 | 121.0 (3) |
| N10—C6—C5 | 125.5 (3) | N20—C16—C15 | 124.9 (3) |
| N7—C8—H8 | 123 (3) | N17—C18—H18 | 125 (3) |
| N9—C8—N7 | 113.6 (3) | N19—C18—N17 | 113.3 (3) |
| N9—C8—H8 | 123 (3) | N19—C18—H18 | 121 (3) |
| | | |
| N3—C4—C5—N7 | −178.7 (3) | N13—C14—C15—N17 | −179.0 (3) |
| N3—C4—C5—C6 | −0.1 (5) | N13—C14—C15—C16 | 2.7 (5) |
| N7—C5—C6—N1 | 178.1 (4) | N17—C15—C16—N11 | −178.8 (4) |
| N7—C5—C6—N10 | −2.2 (7) | N17—C15—C16—N20 | 0.3 (7) |
| N9—C4—C5—N7 | 0.3 (4) | N19—C14—C15—N17 | −0.6 (4) |
| N9—C4—C5—C6 | 178.8 (3) | N19—C14—C15—C16 | −179.0 (3) |
| C2—N1—C6—N10 | 179.4 (3) | C12—N11—C16—N20 | −178.9 (4) |
| C2—N1—C6—C5 | −0.8 (5) | C12—N11—C16—C15 | 0.3 (5) |
| C2—N3—C4—N9 | −178.0 (3) | C12—N13—C14—N19 | 178.5 (3) |
| C2—N3—C4—C5 | 0.8 (5) | C12—N13—C14—C15 | −3.4 (5) |
| C4—N3—C2—O1 | 179.2 (4) | C14—N13—C12—O2 | −178.1 (4) |
| C4—N3—C2—N1 | −1.4 (5) | C14—N13—C12—N11 | 2.4 (5) |
| C4—N9—C8—N7 | −0.7 (4) | C14—N19—C18—N17 | 0.3 (4) |
| C4—C5—C6—N1 | 0.1 (5) | C14—C15—C16—N11 | −1.1 (5) |
| C4—C5—C6—N10 | 179.8 (4) | C14—C15—C16—N20 | 178.0 (4) |
| C5—N7—C8—N9 | 0.9 (4) | C15—N17—C18—N19 | −0.7 (4) |
| C6—N1—C2—O1 | −179.0 (4) | C16—N11—C12—O2 | 179.6 (4) |
| C6—N1—C2—N3 | 1.5 (5) | C16—N11—C12—N13 | −0.9 (5) |
| C8—N7—C5—C4 | −0.7 (4) | C18—N17—C15—C14 | 0.8 (4) |
| C8—N7—C5—C6 | −178.9 (4) | C18—N17—C15—C16 | 178.8 (4) |
| C8—N9—C4—N3 | 179.1 (3) | C18—N19—C14—N13 | 178.5 (4) |
| C8—N9—C4—C5 | 0.2 (4) | C18—N19—C14—C15 | 0.2 (4) |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| N1—H1···Cl1 | 0.89 (6) | 2.18 (6) | 3.067 (3) | 175 (5) |
| N3—H3···Cl1i | 0.95 (5) | 2.13 (5) | 3.078 (3) | 175 (4) |
| N7—H7···Cl2ii | 0.80 (6) | 2.56 (6) | 3.240 (4) | 144 (5) |
| N7—H7···O2iii | 0.80 (6) | 2.48 (6) | 3.014 (4) | 125 (5) |
| N10—H10A···Cl2ii | 0.91 (5) | 2.24 (5) | 3.146 (3) | 174 (4) |
| N10—H10B···N9iv | 0.87 (5) | 2.34 (5) | 3.006 (5) | 134 (4) |
| N11—H11···N19v | 0.90 (6) | 2.07 (6) | 2.957 (4) | 167 (4) |
| N13—H13···Cl2 | 0.85 (4) | 2.25 (5) | 3.093 (3) | 176 (4) |
| N17—H17···Cl1 | 0.86 (4) | 2.47 (4) | 3.267 (3) | 154 (4) |
| N17—H17···O1vi | 0.86 (4) | 2.48 (4) | 2.981 (4) | 118 (4) |
| C18—H18···O1vi | 0.97 (5) | 2.57 (4) | 3.057 (5) | 111 (3) |
| C18—H18···O2vii | 0.97 (5) | 2.26 (5) | 3.078 (4) | 141 (3) |
| N20—H20A···Cl1 | 0.88 (5) | 2.31 (5) | 3.188 (3) | 173 (4) |
| N20—H20B···Cl2v | 0.91 (7) | 2.28 (6) | 3.045 (3) | 141 (5) |
| Symmetry codes: (i) x, y, z−1; (ii) x+1, −y+1/2, z+1/2; (iii) x+1, y, z; (iv) x, y, z+1; (v) −x, y+1/2, −z−1/2; (vi) x, −y+1/2, z+1/2; (vii) −x, y−1/2, −z−1/2. |
Interaction energy (kcal mol-1) of selected dimers in iGH and GH chloride
crystals top | iGHCl | | GUANCD02 | | GUANCH01 | |
| dim1 (tape1) | dim2 (tape2) | dim1 | dim2 | dim1 | dim2 |
| Ees | 43.2 | 31.6 | 41.9 | 40.5 | 8.5 | 42.1 |
| ECoul | 43.1 | 47.7 | 48.0 | 43.2 | 51.0 | 47.6 |
| ECoul-Ees | -0.1 | 16.1 | 6.1 | 2.7 | 42.5 | 5.6 |
| Centre of mass distance (Å) | 7.70 | 6.96 | 6.92 | 7.68 | 6.51 | 6.97 |

 Subscribe to Acta Crystallographica Section C: Structural Chemistry
Subscribe to Acta Crystallographica Section C: Structural Chemistry
The full text of this article is available to subscribers to the journal.
If you have already registered and are using a computer listed in your registration details, please email
support@iucr.org for assistance.
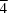 ). The tapes are formed by dimers of isoguaninium cations interacting with chloride anions. Ees analysis indicates that cations in one kind of tape are oriented so as to minimize repulsive electrostatic interactions.
). The tapes are formed by dimers of isoguaninium cations interacting with chloride anions. Ees analysis indicates that cations in one kind of tape are oriented so as to minimize repulsive electrostatic interactions.
 journal menu
journal menu









 Subscribe to Acta Crystallographica Section C: Structural Chemistry
Subscribe to Acta Crystallographica Section C: Structural Chemistry


