Buy article online - an online subscription or single-article purchase is required to access this article.
The crystal structures of 9-(4-vinylbenzyl)adenine, C
14H
13N
5, and 1-(4-vinylbenzyl)uracil, C
13H
12N
2O
2, are composed of zigzag ribbon-like structures that are stabilized by conventional (N—H

N-type) hydrogen bonds for the former and conventional (N—H

O-type) and non-conventional (C—H

O-type) hydrogen bonds for the latter; the hydrogen-bonding patterns are represented by graph-sets
R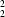
(9) and
R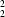
(8), respectively. The adenine and uracil moieties in these alkylated derivatives are planar and are inclined at angles of 84.44 (4) and 79.07 (7)°, respectively, with respect to the phenyl rings.
Supporting information
CCDC references: 192965; 192966
Compounds (I) and (II) were prepared by alkylation of adenine and uracil with
4-vinylbenzyl chloride, using anhydrous carbonate in dimethyl sulfoxide. These
analogs were thoroughly characterized by usual spectroscopic techniques
(Srivatsan & Verma, 2002). Compound (I) was recrystallized from
methanol/chloroform (1:1), while (II) was recrystallized from methanol.
Details of the syntheses and characterization of these derivatives will be
reported elsewhere.
In (I) and (II), most of the H atoms were visible from difference Fourier
syntheses and all were included in the refinements at geometrically idealized
positions, with C—H distances in the range 0.95–0.99 Å and N—H
distances of 0.88 Å, utilizing riding models; H atoms attached to N19 in (I)
were allowed to refine. H atoms were given Uiso values of 1.2 times
the equivalent isotropic displacement parameters of the atoms to which they
were bonded. The absolute structures could not be determined in this analysis.
For both compounds, data collection: COLLECT (Hooft, 1998); cell refinement: HKL DENZO (Otwinowski & Minor, 1997); data reduction: SCALEPACK (Otwinowski & Minor, 1997); program(s) used to solve structure: SAPI91 (Fan, 1991); program(s) used to refine structure: SHELXL97 (Sheldrick, 1997); molecular graphics: ORTEPII (Johnson, 1976); software used to prepare material for publication: SHELXL97.
(I) 9-(4-vinylbenzyl)adenine
top
Crystal data top
| C14H13N5 | Dx = 1.340 Mg m−3 |
| Mr = 251.29 | Mo Kα radiation, λ = 0.71073 Å |
| Orthorhombic, P212121 | Cell parameters from 9411 reflections |
| a = 5.2885 (1) Å | θ = 1.0–30.5° |
| b = 8.1643 (2) Å | µ = 0.09 mm−1 |
| c = 28.8446 (5) Å | T = 170 K |
| V = 1245.42 (4) Å3 | Block, colourless |
| Z = 4 | 0.25 × 0.22 × 0.20 mm |
| F(000) = 528 | |
Data collection top
Nonius KappaCCD
diffractometer | 2201 independent reflections |
| Radiation source: fine-focus sealed tube | 1988 reflections with I > 2σ(I) |
| Graphite monochromator | Rint = 0.052 |
| ω and ϕ scans | θmax = 30.5°, θmin = 3.7° |
Absorption correction: multi-scan
(SORTAV: Blessing, 1995, 1997) | h = −6→7 |
| Tmin = 0.979, Tmax = 0.983 | k = −11→11 |
| 9411 measured reflections | l = −40→41 |
Refinement top
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.039 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.111 | H-atom parameters constrained |
| S = 1.04 | w = 1/[σ2(Fo2) + (0.0774P)2 + 0.0463P]
where P = (Fo2 + 2Fc2)/3 |
| 2201 reflections | (Δ/σ)max < 0.001 |
| 178 parameters | Δρmax = 0.27 e Å−3 |
| 0 restraints | Δρmin = −0.22 e Å−3 |
Crystal data top
| C14H13N5 | V = 1245.42 (4) Å3 |
| Mr = 251.29 | Z = 4 |
| Orthorhombic, P212121 | Mo Kα radiation |
| a = 5.2885 (1) Å | µ = 0.09 mm−1 |
| b = 8.1643 (2) Å | T = 170 K |
| c = 28.8446 (5) Å | 0.25 × 0.22 × 0.20 mm |
Data collection top
Nonius KappaCCD
diffractometer | 2201 independent reflections |
Absorption correction: multi-scan
(SORTAV: Blessing, 1995, 1997) | 1988 reflections with I > 2σ(I) |
| Tmin = 0.979, Tmax = 0.983 | Rint = 0.052 |
| 9411 measured reflections | |
Refinement top
| R[F2 > 2σ(F2)] = 0.039 | 0 restraints |
| wR(F2) = 0.111 | H-atom parameters constrained |
| S = 1.04 | Δρmax = 0.27 e Å−3 |
| 2201 reflections | Δρmin = −0.22 e Å−3 |
| 178 parameters | |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| N1 | 0.7308 (3) | 0.35469 (15) | 0.22489 (4) | 0.0294 (3) | |
| C2 | 0.5350 (3) | 0.39463 (18) | 0.19748 (5) | 0.0317 (3) | |
| H2 | 0.4912 | 0.5075 | 0.1973 | 0.038* | |
| N3 | 0.3936 (3) | 0.29980 (16) | 0.17058 (4) | 0.0321 (3) | |
| C4 | 0.4712 (3) | 0.14274 (17) | 0.17367 (5) | 0.0266 (3) | |
| C5 | 0.6706 (3) | 0.08190 (17) | 0.20015 (4) | 0.0254 (3) | |
| C6 | 0.8046 (3) | 0.19602 (17) | 0.22750 (4) | 0.0260 (3) | |
| N7 | 0.6933 (3) | −0.08633 (15) | 0.19380 (4) | 0.0286 (3) | |
| C8 | 0.5115 (3) | −0.12184 (18) | 0.16470 (5) | 0.0298 (3) | |
| H8 | 0.4802 | −0.2299 | 0.1539 | 0.036* | |
| N9 | 0.3706 (3) | 0.00982 (15) | 0.15116 (4) | 0.0293 (3) | |
| C10 | 0.1710 (3) | 0.0130 (2) | 0.11618 (5) | 0.0320 (3) | |
| H10A | 0.0679 | 0.1131 | 0.1205 | 0.038* | |
| H10B | 0.0590 | −0.0828 | 0.1207 | 0.038* | |
| C11 | 0.2737 (3) | 0.01014 (18) | 0.06718 (5) | 0.0263 (3) | |
| C12 | 0.4913 (3) | 0.0964 (2) | 0.05489 (5) | 0.0303 (3) | |
| H12 | 0.5830 | 0.1550 | 0.0779 | 0.036* | |
| C13 | 0.1441 (3) | −0.0759 (2) | 0.03292 (5) | 0.0305 (3) | |
| H13 | −0.0022 | −0.1373 | 0.0409 | 0.037* | |
| C14 | 0.5745 (3) | 0.0972 (2) | 0.00921 (5) | 0.0313 (3) | |
| H14 | 0.7241 | 0.1555 | 0.0015 | 0.038* | |
| C15 | 0.2262 (3) | −0.0730 (2) | −0.01278 (5) | 0.0321 (3) | |
| H15 | 0.1338 | −0.1311 | −0.0358 | 0.038* | |
| C16 | 0.4433 (3) | 0.01417 (18) | −0.02551 (5) | 0.0286 (3) | |
| C17 | 0.5366 (3) | 0.0235 (2) | −0.07373 (5) | 0.0361 (4) | |
| H17 | 0.6945 | 0.0767 | −0.0781 | 0.043* | |
| C18 | 0.4268 (5) | −0.0329 (3) | −0.11093 (6) | 0.0618 (6) | |
| H18A | 0.2685 | −0.0872 | −0.1086 | 0.074* | |
| H18B | 0.5052 | −0.0196 | −0.1403 | 0.074* | |
| N19 | 0.9936 (3) | 0.15772 (16) | 0.25591 (5) | 0.0318 (3) | |
| H19A | 1.091 (4) | 0.245 (2) | 0.2703 (7) | 0.038* | |
| H19B | 1.058 (4) | 0.048 (3) | 0.2561 (7) | 0.038* | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| N1 | 0.0368 (7) | 0.0221 (5) | 0.0292 (6) | 0.0017 (5) | 0.0005 (5) | 0.0010 (4) |
| C2 | 0.0400 (8) | 0.0242 (6) | 0.0309 (7) | 0.0046 (6) | 0.0015 (6) | 0.0028 (5) |
| N3 | 0.0373 (7) | 0.0287 (6) | 0.0303 (6) | 0.0053 (6) | −0.0005 (5) | 0.0024 (5) |
| C4 | 0.0303 (7) | 0.0281 (6) | 0.0215 (5) | 0.0013 (6) | 0.0026 (5) | 0.0008 (5) |
| C5 | 0.0309 (7) | 0.0229 (6) | 0.0222 (5) | 0.0003 (6) | 0.0020 (5) | 0.0015 (5) |
| C6 | 0.0314 (7) | 0.0238 (6) | 0.0227 (6) | −0.0010 (6) | 0.0030 (5) | 0.0022 (5) |
| N7 | 0.0353 (6) | 0.0228 (5) | 0.0276 (5) | −0.0004 (5) | 0.0001 (5) | 0.0007 (4) |
| C8 | 0.0370 (8) | 0.0254 (6) | 0.0271 (6) | −0.0017 (6) | 0.0010 (6) | 0.0005 (5) |
| N9 | 0.0324 (6) | 0.0301 (6) | 0.0255 (5) | 0.0001 (6) | −0.0015 (5) | 0.0008 (5) |
| C10 | 0.0262 (6) | 0.0421 (8) | 0.0279 (6) | −0.0024 (7) | −0.0009 (5) | 0.0003 (6) |
| C11 | 0.0243 (6) | 0.0278 (6) | 0.0268 (6) | 0.0008 (6) | −0.0011 (5) | 0.0013 (5) |
| C12 | 0.0262 (6) | 0.0351 (7) | 0.0296 (6) | −0.0051 (6) | −0.0030 (5) | −0.0009 (6) |
| C13 | 0.0269 (7) | 0.0314 (7) | 0.0331 (7) | −0.0037 (6) | −0.0012 (6) | −0.0013 (6) |
| C14 | 0.0257 (7) | 0.0345 (7) | 0.0336 (7) | −0.0030 (6) | 0.0007 (5) | 0.0018 (6) |
| C15 | 0.0323 (7) | 0.0333 (7) | 0.0307 (7) | −0.0020 (7) | −0.0042 (6) | −0.0048 (6) |
| C16 | 0.0298 (7) | 0.0283 (7) | 0.0277 (6) | 0.0041 (6) | −0.0002 (5) | −0.0011 (5) |
| C17 | 0.0379 (8) | 0.0387 (8) | 0.0318 (7) | 0.0019 (7) | 0.0044 (6) | 0.0009 (6) |
| C18 | 0.0686 (14) | 0.0886 (17) | 0.0283 (7) | −0.0208 (14) | −0.0002 (8) | 0.0012 (9) |
| N19 | 0.0392 (7) | 0.0222 (5) | 0.0340 (6) | −0.0012 (6) | −0.0083 (6) | −0.0006 (5) |
Geometric parameters (Å, º) top
| N1—C2 | 1.343 (2) | C11—C13 | 1.393 (2) |
| N1—C6 | 1.3550 (17) | C11—C12 | 1.395 (2) |
| C2—N3 | 1.327 (2) | C12—C14 | 1.389 (2) |
| C2—H2 | 0.9500 | C12—H12 | 0.9500 |
| N3—C4 | 1.3494 (18) | C13—C15 | 1.388 (2) |
| C4—N9 | 1.3720 (19) | C13—H13 | 0.9500 |
| C4—C5 | 1.393 (2) | C14—C16 | 1.394 (2) |
| C5—N7 | 1.3909 (18) | C14—H14 | 0.9500 |
| C5—C6 | 1.412 (2) | C15—C16 | 1.400 (2) |
| C6—N19 | 1.330 (2) | C15—H15 | 0.9500 |
| N7—C8 | 1.309 (2) | C16—C17 | 1.4777 (19) |
| C8—N9 | 1.3648 (19) | C17—C18 | 1.304 (3) |
| C8—H8 | 0.9500 | C17—H17 | 0.9500 |
| N9—C10 | 1.4608 (19) | C18—H18A | 0.9500 |
| C10—C11 | 1.5144 (19) | C18—H18B | 0.9500 |
| C10—H10A | 0.9900 | N19—H19A | 0.97 (2) |
| C10—H10B | 0.9900 | N19—H19B | 0.96 (2) |
| | | |
| C2—N1—C6 | 119.13 (13) | C13—C11—C12 | 118.71 (13) |
| N3—C2—N1 | 129.58 (14) | C13—C11—C10 | 119.57 (13) |
| N3—C2—H2 | 115.2 | C12—C11—C10 | 121.68 (13) |
| N1—C2—H2 | 115.2 | C14—C12—C11 | 120.28 (14) |
| C2—N3—C4 | 110.13 (13) | C14—C12—H12 | 119.9 |
| N3—C4—N9 | 127.04 (14) | C11—C12—H12 | 119.9 |
| N3—C4—C5 | 127.25 (14) | C15—C13—C11 | 120.74 (14) |
| N9—C4—C5 | 105.71 (12) | C15—C13—H13 | 119.6 |
| N7—C5—C4 | 110.20 (13) | C11—C13—H13 | 119.6 |
| N7—C5—C6 | 133.00 (14) | C12—C14—C16 | 121.45 (14) |
| C4—C5—C6 | 116.80 (13) | C12—C14—H14 | 119.3 |
| N19—C6—N1 | 118.39 (13) | C16—C14—H14 | 119.3 |
| N19—C6—C5 | 124.51 (13) | C13—C15—C16 | 120.96 (14) |
| N1—C6—C5 | 117.09 (13) | C13—C15—H15 | 119.5 |
| C8—N7—C5 | 103.86 (13) | C16—C15—H15 | 119.5 |
| N7—C8—N9 | 114.21 (13) | C14—C16—C15 | 117.84 (13) |
| N7—C8—H8 | 122.9 | C14—C16—C17 | 119.00 (14) |
| N9—C8—H8 | 122.9 | C15—C16—C17 | 123.16 (14) |
| C8—N9—C4 | 106.02 (12) | C18—C17—C16 | 127.42 (18) |
| C8—N9—C10 | 127.31 (13) | C18—C17—H17 | 116.3 |
| C4—N9—C10 | 126.38 (13) | C16—C17—H17 | 116.3 |
| N9—C10—C11 | 112.66 (12) | C17—C18—H18A | 120.0 |
| N9—C10—H10A | 109.1 | C17—C18—H18B | 120.0 |
| C11—C10—H10A | 109.1 | H18A—C18—H18B | 120.0 |
| N9—C10—H10B | 109.1 | C6—N19—H19A | 119.3 (12) |
| C11—C10—H10B | 109.1 | C6—N19—H19B | 119.4 (12) |
| H10A—C10—H10B | 107.8 | H19A—N19—H19B | 119.6 (17) |
| | | |
| C6—N1—C2—N3 | −0.6 (2) | C5—C4—N9—C8 | −0.20 (16) |
| N1—C2—N3—C4 | 0.3 (2) | N3—C4—N9—C10 | 5.4 (2) |
| C2—N3—C4—N9 | 179.68 (14) | C5—C4—N9—C10 | −174.36 (13) |
| C2—N3—C4—C5 | −0.7 (2) | C8—N9—C10—C11 | −76.27 (19) |
| N3—C4—C5—N7 | −179.37 (14) | C4—N9—C10—C11 | 96.67 (17) |
| N9—C4—C5—N7 | 0.34 (16) | N9—C10—C11—C13 | 143.69 (14) |
| N3—C4—C5—C6 | 1.2 (2) | N9—C10—C11—C12 | −38.5 (2) |
| N9—C4—C5—C6 | −179.11 (12) | C13—C11—C12—C14 | 0.6 (2) |
| C2—N1—C6—N19 | −177.91 (14) | C10—C11—C12—C14 | −177.18 (14) |
| C2—N1—C6—C5 | 1.0 (2) | C12—C11—C13—C15 | −1.4 (2) |
| N7—C5—C6—N19 | −1.7 (3) | C10—C11—C13—C15 | 176.39 (15) |
| C4—C5—C6—N19 | 177.58 (14) | C11—C12—C14—C16 | 0.7 (2) |
| N7—C5—C6—N1 | 179.45 (14) | C11—C13—C15—C16 | 1.0 (2) |
| C4—C5—C6—N1 | −1.25 (19) | C12—C14—C16—C15 | −1.2 (2) |
| C4—C5—N7—C8 | −0.34 (16) | C12—C14—C16—C17 | 178.14 (15) |
| C6—C5—N7—C8 | 178.99 (16) | C13—C15—C16—C14 | 0.3 (2) |
| C5—N7—C8—N9 | 0.21 (17) | C13—C15—C16—C17 | −178.96 (15) |
| N7—C8—N9—C4 | −0.01 (18) | C14—C16—C17—C18 | −173.2 (2) |
| N7—C8—N9—C10 | 174.08 (13) | C15—C16—C17—C18 | 6.1 (3) |
| N3—C4—N9—C8 | 179.51 (15) | | |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| N19—H19A···N7i | 0.97 (2) | 2.07 (2) | 3.0351 (19) | 174 (2) |
| N19—H19B···N1ii | 0.96 (2) | 2.01 (2) | 2.9242 (19) | 159 (2) |
| Symmetry codes: (i) −x+2, y+1/2, −z+1/2; (ii) −x+2, y−1/2, −z+1/2. |
(II) 1-(4-vinylbenzyl)uracil
top
Crystal data top
| C13H12N2O2 | Dx = 1.395 Mg m−3 |
| Mr = 228.25 | Mo Kα radiation, λ = 0.71073 Å |
| Orthorhombic, P212121 | Cell parameters from 6507 reflections |
| a = 5.9603 (2) Å | θ = 1.0–30.6° |
| b = 7.0132 (2) Å | µ = 0.10 mm−1 |
| c = 25.9571 (9) Å | T = 170 K |
| V = 1085.03 (6) Å3 | Prism, colourless |
| Z = 4 | 0.18 × 0.15 × 0.14 mm |
| F(000) = 480 | |
Data collection top
Nonius KappaCCD
diffractometer | 1901 independent reflections |
| Radiation source: fine-focus sealed tube | 1525 reflections with I > 2σ(I) |
| Graphite monochromator | Rint = 0.098 |
| ω and ϕ scans | θmax = 30.6°, θmin = 3.7° |
Absorption correction: multi-scan
(SORTAV: Blessing, 1995, 1997) | h = −8→8 |
| Tmin = 0.983, Tmax = 0.986 | k = −5→10 |
| 6507 measured reflections | l = −35→37 |
Refinement top
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.054 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.135 | H-atom parameters constrained |
| S = 1.00 | w = 1/[σ2(Fo2) + (0.0658P)2 + 0.3792P]
where P = (Fo2 + 2Fc2)/3 |
| 1901 reflections | (Δ/σ)max < 0.001 |
| 154 parameters | Δρmax = 0.31 e Å−3 |
| 0 restraints | Δρmin = −0.32 e Å−3 |
Crystal data top
| C13H12N2O2 | V = 1085.03 (6) Å3 |
| Mr = 228.25 | Z = 4 |
| Orthorhombic, P212121 | Mo Kα radiation |
| a = 5.9603 (2) Å | µ = 0.10 mm−1 |
| b = 7.0132 (2) Å | T = 170 K |
| c = 25.9571 (9) Å | 0.18 × 0.15 × 0.14 mm |
Data collection top
Nonius KappaCCD
diffractometer | 1901 independent reflections |
Absorption correction: multi-scan
(SORTAV: Blessing, 1995, 1997) | 1525 reflections with I > 2σ(I) |
| Tmin = 0.983, Tmax = 0.986 | Rint = 0.098 |
| 6507 measured reflections | |
Refinement top
| R[F2 > 2σ(F2)] = 0.054 | 0 restraints |
| wR(F2) = 0.135 | H-atom parameters constrained |
| S = 1.00 | Δρmax = 0.31 e Å−3 |
| 1901 reflections | Δρmin = −0.32 e Å−3 |
| 154 parameters | |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| O1 | 0.0003 (3) | 0.3922 (3) | 0.24879 (7) | 0.0292 (4) | |
| O2 | 0.5017 (3) | 0.7921 (2) | 0.32375 (6) | 0.0282 (4) | |
| N1 | 0.5612 (3) | 0.4711 (3) | 0.33265 (7) | 0.0223 (4) | |
| C2 | 0.4449 (4) | 0.6285 (3) | 0.31491 (8) | 0.0221 (5) | |
| N3 | 0.2548 (3) | 0.5856 (3) | 0.28640 (7) | 0.0225 (4) | |
| H3 | 0.1766 | 0.6828 | 0.2747 | 0.027* | |
| C4 | 0.1757 (4) | 0.4059 (4) | 0.27443 (8) | 0.0218 (5) | |
| C5 | 0.3071 (4) | 0.2493 (4) | 0.29408 (8) | 0.0239 (5) | |
| H5 | 0.2644 | 0.1212 | 0.2874 | 0.029* | |
| C6 | 0.4905 (4) | 0.2884 (3) | 0.32191 (8) | 0.0242 (5) | |
| H6 | 0.5762 | 0.1849 | 0.3349 | 0.029* | |
| C7 | 0.7642 (4) | 0.5008 (4) | 0.36394 (8) | 0.0251 (5) | |
| H7A | 0.8324 | 0.6246 | 0.3544 | 0.030* | |
| H7B | 0.8741 | 0.3995 | 0.3557 | 0.030* | |
| C8 | 0.7199 (4) | 0.4996 (3) | 0.42150 (8) | 0.0218 (4) | |
| C9 | 0.5221 (4) | 0.5691 (4) | 0.44304 (8) | 0.0249 (5) | |
| H9 | 0.4058 | 0.6144 | 0.4212 | 0.030* | |
| C10 | 0.8878 (4) | 0.4340 (4) | 0.45403 (9) | 0.0252 (5) | |
| H10 | 1.0238 | 0.3866 | 0.4399 | 0.030* | |
| C11 | 0.4935 (4) | 0.5727 (4) | 0.49629 (8) | 0.0251 (5) | |
| H11 | 0.3576 | 0.6207 | 0.5104 | 0.030* | |
| C12 | 0.8585 (4) | 0.4372 (4) | 0.50729 (9) | 0.0253 (5) | |
| H12 | 0.9748 | 0.3911 | 0.5290 | 0.030* | |
| C13 | 0.6610 (4) | 0.5071 (4) | 0.52922 (8) | 0.0235 (5) | |
| C14 | 0.6372 (5) | 0.5091 (4) | 0.58588 (8) | 0.0281 (5) | |
| H14 | 0.7706 | 0.4929 | 0.6054 | 0.034* | |
| C15 | 0.4486 (5) | 0.5313 (4) | 0.61217 (9) | 0.0351 (6) | |
| H15A | 0.3106 | 0.5481 | 0.5944 | 0.042* | |
| H15B | 0.4514 | 0.5305 | 0.6488 | 0.042* | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| O1 | 0.0293 (8) | 0.0285 (9) | 0.0297 (7) | −0.0018 (8) | −0.0100 (7) | −0.0011 (7) |
| O2 | 0.0324 (10) | 0.0266 (8) | 0.0257 (8) | −0.0033 (8) | −0.0031 (8) | −0.0013 (7) |
| N1 | 0.0212 (9) | 0.0294 (11) | 0.0163 (7) | −0.0002 (8) | −0.0017 (7) | 0.0002 (7) |
| C2 | 0.0232 (11) | 0.0281 (11) | 0.0151 (8) | −0.0007 (9) | 0.0005 (8) | −0.0002 (8) |
| N3 | 0.0245 (10) | 0.0232 (9) | 0.0198 (8) | −0.0003 (9) | −0.0025 (8) | 0.0007 (7) |
| C4 | 0.0246 (11) | 0.0259 (11) | 0.0150 (8) | −0.0022 (10) | 0.0018 (8) | 0.0005 (8) |
| C5 | 0.0298 (12) | 0.0226 (10) | 0.0193 (9) | −0.0001 (10) | 0.0015 (10) | −0.0006 (8) |
| C6 | 0.0262 (13) | 0.0263 (11) | 0.0200 (9) | 0.0020 (10) | 0.0011 (10) | 0.0016 (9) |
| C7 | 0.0188 (10) | 0.0379 (13) | 0.0187 (9) | 0.0015 (10) | −0.0011 (8) | −0.0004 (10) |
| C8 | 0.0234 (11) | 0.0242 (10) | 0.0177 (8) | −0.0004 (9) | −0.0020 (8) | 0.0001 (9) |
| C9 | 0.0220 (11) | 0.0326 (11) | 0.0200 (9) | 0.0038 (11) | −0.0039 (9) | 0.0025 (9) |
| C10 | 0.0221 (11) | 0.0301 (12) | 0.0234 (10) | 0.0038 (10) | −0.0015 (8) | −0.0017 (9) |
| C11 | 0.0251 (11) | 0.0293 (11) | 0.0209 (9) | 0.0040 (11) | 0.0008 (9) | 0.0000 (9) |
| C12 | 0.0254 (11) | 0.0277 (11) | 0.0229 (9) | 0.0039 (10) | −0.0071 (9) | 0.0001 (9) |
| C13 | 0.0284 (11) | 0.0237 (10) | 0.0183 (8) | −0.0012 (10) | −0.0013 (9) | −0.0005 (9) |
| C14 | 0.0355 (13) | 0.0313 (13) | 0.0174 (9) | −0.0011 (12) | −0.0041 (9) | 0.0009 (10) |
| C15 | 0.0457 (15) | 0.0397 (15) | 0.0200 (10) | 0.0022 (13) | 0.0029 (10) | 0.0003 (10) |
Geometric parameters (Å, º) top
| O1—C4 | 1.243 (3) | C8—C10 | 1.388 (3) |
| O2—C2 | 1.218 (3) | C8—C9 | 1.393 (3) |
| N1—C6 | 1.377 (3) | C9—C11 | 1.393 (3) |
| N1—C2 | 1.382 (3) | C9—H9 | 0.9500 |
| N1—C7 | 1.472 (3) | C10—C12 | 1.394 (3) |
| C2—N3 | 1.386 (3) | C10—H10 | 0.9500 |
| N3—C4 | 1.381 (3) | C11—C13 | 1.393 (3) |
| N3—H3 | 0.8800 | C11—H11 | 0.9500 |
| C4—C5 | 1.442 (3) | C12—C13 | 1.396 (3) |
| C5—C6 | 1.338 (3) | C12—H12 | 0.9500 |
| C5—H5 | 0.9500 | C13—C14 | 1.478 (3) |
| C6—H6 | 0.9500 | C14—C15 | 1.324 (4) |
| C7—C8 | 1.517 (3) | C14—H14 | 0.9500 |
| C7—H7A | 0.9900 | C15—H15A | 0.9500 |
| C7—H7B | 0.9900 | C15—H15B | 0.9500 |
| | | |
| C6—N1—C2 | 121.47 (19) | C10—C8—C9 | 118.81 (19) |
| C6—N1—C7 | 119.7 (2) | C10—C8—C7 | 118.4 (2) |
| C2—N1—C7 | 118.9 (2) | C9—C8—C7 | 122.7 (2) |
| O2—C2—N1 | 123.4 (2) | C8—C9—C11 | 120.6 (2) |
| O2—C2—N3 | 122.1 (2) | C8—C9—H9 | 119.7 |
| N1—C2—N3 | 114.5 (2) | C11—C9—H9 | 119.7 |
| C4—N3—C2 | 126.7 (2) | C8—C10—C12 | 120.6 (2) |
| C4—N3—H3 | 116.7 | C8—C10—H10 | 119.7 |
| C2—N3—H3 | 116.7 | C12—C10—H10 | 119.7 |
| O1—C4—N3 | 118.6 (2) | C13—C11—C9 | 121.0 (2) |
| O1—C4—C5 | 126.0 (2) | C13—C11—H11 | 119.5 |
| N3—C4—C5 | 115.5 (2) | C9—C11—H11 | 119.5 |
| C6—C5—C4 | 118.6 (2) | C10—C12—C13 | 121.0 (2) |
| C6—C5—H5 | 120.7 | C10—C12—H12 | 119.5 |
| C4—C5—H5 | 120.7 | C13—C12—H12 | 119.5 |
| C5—C6—N1 | 123.3 (2) | C11—C13—C12 | 118.05 (19) |
| C5—C6—H6 | 118.3 | C11—C13—C14 | 122.6 (2) |
| N1—C6—H6 | 118.3 | C12—C13—C14 | 119.4 (2) |
| N1—C7—C8 | 113.53 (18) | C15—C14—C13 | 126.6 (2) |
| N1—C7—H7A | 108.9 | C15—C14—H14 | 116.7 |
| C8—C7—H7A | 108.9 | C13—C14—H14 | 116.7 |
| N1—C7—H7B | 108.9 | C14—C15—H15A | 120.0 |
| C8—C7—H7B | 108.9 | C14—C15—H15B | 120.0 |
| H7A—C7—H7B | 107.7 | H15A—C15—H15B | 120.0 |
| | | |
| C6—N1—C2—O2 | 179.7 (2) | N1—C7—C8—C10 | 149.0 (2) |
| C7—N1—C2—O2 | 0.1 (3) | N1—C7—C8—C9 | −33.9 (4) |
| C6—N1—C2—N3 | 0.4 (3) | C10—C8—C9—C11 | 0.0 (4) |
| C7—N1—C2—N3 | −179.22 (18) | C7—C8—C9—C11 | −177.1 (2) |
| O2—C2—N3—C4 | −179.7 (2) | C9—C8—C10—C12 | 0.2 (4) |
| N1—C2—N3—C4 | −0.4 (3) | C7—C8—C10—C12 | 177.4 (2) |
| C2—N3—C4—O1 | 179.2 (2) | C8—C9—C11—C13 | −0.1 (4) |
| C2—N3—C4—C5 | 0.0 (3) | C8—C10—C12—C13 | −0.4 (4) |
| O1—C4—C5—C6 | −178.8 (2) | C9—C11—C13—C12 | −0.1 (4) |
| N3—C4—C5—C6 | 0.3 (3) | C9—C11—C13—C14 | 179.8 (2) |
| C4—C5—C6—N1 | −0.3 (3) | C10—C12—C13—C11 | 0.3 (4) |
| C2—N1—C6—C5 | −0.1 (3) | C10—C12—C13—C14 | −179.6 (2) |
| C7—N1—C6—C5 | 179.54 (19) | C11—C13—C14—C15 | 15.5 (5) |
| C6—N1—C7—C8 | −85.9 (3) | C12—C13—C14—C15 | −164.6 (3) |
| C2—N1—C7—C8 | 93.8 (3) | | |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| N3—H3···O1i | 0.88 | 1.91 | 2.788 (3) | 178 |
| C5—H5···O1ii | 0.95 | 2.44 | 3.296 (3) | 150 |
| Symmetry codes: (i) −x, y+1/2, −z+1/2; (ii) −x, y−1/2, −z+1/2. |
Experimental details
| | (I) | (II) |
| Crystal data |
| Chemical formula | C14H13N5 | C13H12N2O2 |
| Mr | 251.29 | 228.25 |
| Crystal system, space group | Orthorhombic, P212121 | Orthorhombic, P212121 |
| Temperature (K) | 170 | 170 |
| a, b, c (Å) | 5.2885 (1), 8.1643 (2), 28.8446 (5) | 5.9603 (2), 7.0132 (2), 25.9571 (9) |
| V (Å3) | 1245.42 (4) | 1085.03 (6) |
| Z | 4 | 4 |
| Radiation type | Mo Kα | Mo Kα |
| µ (mm−1) | 0.09 | 0.10 |
| Crystal size (mm) | 0.25 × 0.22 × 0.20 | 0.18 × 0.15 × 0.14 |
| |
| Data collection |
| Diffractometer | Nonius KappaCCD
diffractometer | Nonius KappaCCD
diffractometer |
| Absorption correction | Multi-scan
(SORTAV: Blessing, 1995, 1997) | Multi-scan
(SORTAV: Blessing, 1995, 1997) |
| Tmin, Tmax | 0.979, 0.983 | 0.983, 0.986 |
No. of measured, independent and
observed [I > 2σ(I)] reflections | 9411, 2201, 1988 | 6507, 1901, 1525 |
| Rint | 0.052 | 0.098 |
| (sin θ/λ)max (Å−1) | 0.714 | 0.716 |
| |
| Refinement |
| R[F2 > 2σ(F2)], wR(F2), S | 0.039, 0.111, 1.04 | 0.054, 0.135, 1.00 |
| No. of reflections | 2201 | 1901 |
| No. of parameters | 178 | 154 |
| H-atom treatment | H-atom parameters constrained | H-atom parameters constrained |
| Δρmax, Δρmin (e Å−3) | 0.27, −0.22 | 0.31, −0.32 |
Selected geometric parameters (Å, º) for (I) top| N1—C2 | 1.343 (2) | C5—N7 | 1.3909 (18) |
| N1—C6 | 1.3550 (17) | C6—N19 | 1.330 (2) |
| C2—N3 | 1.327 (2) | C8—N9 | 1.3648 (19) |
| N3—C4 | 1.3494 (18) | N9—C10 | 1.4608 (19) |
| C4—N9 | 1.3720 (19) | C17—C18 | 1.304 (3) |
| | | |
| C2—N1—C6 | 119.13 (13) | C8—N9—C4 | 106.02 (12) |
| C2—N3—C4 | 110.13 (13) | C8—N9—C10 | 127.31 (13) |
| C8—N7—C5 | 103.86 (13) | C4—N9—C10 | 126.38 (13) |
Hydrogen-bond geometry (Å, º) for (I) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| N19—H19A···N7i | 0.97 (2) | 2.07 (2) | 3.0351 (19) | 174 (2) |
| N19—H19B···N1ii | 0.96 (2) | 2.01 (2) | 2.9242 (19) | 159 (2) |
| Symmetry codes: (i) −x+2, y+1/2, −z+1/2; (ii) −x+2, y−1/2, −z+1/2. |
Selected geometric parameters (Å, º) for (II) top| O1—C4 | 1.243 (3) | N1—C7 | 1.472 (3) |
| O2—C2 | 1.218 (3) | N3—C4 | 1.381 (3) |
| N1—C6 | 1.377 (3) | C14—C15 | 1.324 (4) |
| N1—C2 | 1.382 (3) | | |
| | | |
| C6—N1—C2 | 121.47 (19) | C2—N1—C7 | 118.9 (2) |
| C6—N1—C7 | 119.7 (2) | C4—N3—C2 | 126.7 (2) |
Hydrogen-bond geometry (Å, º) for (II) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| N3—H3···O1i | 0.88 | 1.91 | 2.788 (3) | 178 |
| C5—H5···O1ii | 0.95 | 2.44 | 3.296 (3) | 150 |
| Symmetry codes: (i) −x, y+1/2, −z+1/2; (ii) −x, y−1/2, −z+1/2. |

 Subscribe to Acta Crystallographica Section C: Structural Chemistry
Subscribe to Acta Crystallographica Section C: Structural Chemistry
The full text of this article is available to subscribers to the journal.
If you have already registered and are using a computer listed in your registration details, please email
support@iucr.org for assistance.
 N-type) hydrogen bonds for the former and conventional (N—H
N-type) hydrogen bonds for the former and conventional (N—H O-type) and non-conventional (C—H
O-type) and non-conventional (C—H O-type) hydrogen bonds for the latter; the hydrogen-bonding patterns are represented by graph-sets R
O-type) hydrogen bonds for the latter; the hydrogen-bonding patterns are represented by graph-sets R
 Subscribe to Acta Crystallographica Section C: Structural Chemistry
Subscribe to Acta Crystallographica Section C: Structural Chemistry
 journal menu
journal menu








![[Figure 1]](http://journals.iucr.org/c/issues/2002/07/00/fr1366/fr1366fig1thm.gif)
![[Figure 2]](http://journals.iucr.org/c/issues/2002/07/00/fr1366/fr1366fig2thm.gif)
![[Figure 3]](http://journals.iucr.org/c/issues/2002/07/00/fr1366/fr1366fig3thm.gif)
![[Figure 4]](http://journals.iucr.org/c/issues/2002/07/00/fr1366/fr1366fig4thm.gif)

 Subscribe to Acta Crystallographica Section C: Structural Chemistry
Subscribe to Acta Crystallographica Section C: Structural Chemistry



Our current research efforts are focused towards developing metallated nucleobase polymeric resins possessing catalytic activity towards phosphate ester hydrolysis (Srivatsan & Verma, 2000, 2001; Madhavaiah et al., 2002). These resins are generally prepared by AIBN-initiated free-radical polymerization of functional monomers such as 9-allyladenine followed by metallation by a copper salt. We have performed extensive kinetic analyses of phosphate ester hydrolysis using copper-metallated nucleobase resins as catalysts and have found significant rate acceleration of hydrolytic reactions over the uncatalyzed rate, using model substrates. We report the preparation of two new reactive nucleobase monomers, 9-(4-vinylbenzyl)adenine, (I), and 1-(4-vinylbenzyl)uracil, (II), for incorporation in polymeric resins to design multiple usage catalysts for phosphate ester hydrolysis. X-Ray crystallographic studies were performed to unequivocally establish the site of alkylation. The structures of (I) and (II) are presented in this paper.
Compound (I), the vinylated adenine monomer wherein a vinylbenzyl group is substituted atthe N9 position of adenine (Fig. 1), utilizes both Watson–Crick and Hoogsteen hydrogen-bonding faces (Saenger, 1984), with the participation of two exocyclic amino H atoms, with inequivalent H···N distances of 2.11 and 2.20 Å, and almost linear N—H···N angles, resulting in a zigzag chain structure that is stabilized by conventional hydrogen bonds (Fig. 2). The hydrogen-bonding pattern in the polymeric motif can be best described by graph-set R22(9) (Etter et al., 1990). The molecular dimensions in (I) are normal and lie within expected values for corresponding bond distances and angles (Orpen et al., 1994). The adenine moiety is almost planar [maximum deviation of 0.0260 (10) Å for N19] and is inclined at an angle 84.44 (4)° to the mean plane of the phenyl ring. The vinyl group is almost coplanar with the phenyl ring; the mean planes of the two groups are inclined at an angle of 6.8 (3)° with respect to one another.
In compound (II), wherein alkylation has taken place at the expected N1 position of uracil (Fig. 3), the H atom attached to C5 in the pyrimidine ring acts as a donor to form a weak hydrogen bond with a carbonyl (O1) acceptor. This O atom behaves as a double acceptor forming a H-bond with an N—H donor as well, thus aiding in the formation of an extended ribbon-like structure that derives stability both from conventional and non-conventional hydrogen bonds (Fig. 4). Details of hydrogen-bonding geometry are given in Table 4. The hydrogen-bonding pattern in the polymeric motif of (II) can also be described by graph-set R22(8) (Etter et al., 1990). A similar structure has been reported for uracil (Stewart & Jensen, 1967), based on a similar hydrogen-bonding pattern. The molecular dimensions in (II) are also normal and lie within expected ranges. The uracil moiety in (II) is essentially planar [maximum deviation of 0.0085 (12) Å] and is oriented at an angle of 79.07 (7)° with respect to the phenyl ring. However, the vinyl group in (II) is not coplanar with the phenyl ring; the mean planes of the two groups are inclined at an angle of 15.4 (3)° with respect to one another.
The formation and co-operativity of hydrogen bonds dictate the structural stability of biological macromolecules such as nucleic acids and proteins (Saenger, 1984). These interactions are responsible for ensuring precise biomolecular recognition and are utilized extensively for stabilization of protein and ribonucleic acid folding events (Rose & Wolfenden, 1993; Moore, 1999). Recently, non-conventional hydrogen-bonding interactions have received significant recognition owing to their occurrence and contribution to the overall stability of biological systems (Wahl & Sundaralingam, 1997; Desiraju & Steiner, 1999). The two reactive nucleobase monomers reported in this paper display both conventional and non-conventional hydrogen bonds in their respective crystal structures, and these interactions have been invoked to stabilize an extended array of modified nucleobases. We intend to harness observed hydrogen-bonding interactions in (I) and (II) to engineer specificity in our polymeric constructs for enhancing substrate–catalyst recognition.