Buy article online - an online subscription or single-article purchase is required to access this article.

In order to extend the experimental data already available on the close packing of cyclopentanes substituted with vicinal CO
X (
X = OH, NH
2) and OH groups to the analogous cyclohexanes, cycloheptanes and cyclooctanes, (1
R*,2
S*)
-cis-2-hydroxy-1-cyclooctanecarboxylic acid (8C), (1
R*,2
R*)-
trans-2-hydroxy-1-cyclooctanecarboxylic acid (8T) and (1
R*,2
R*)-
trans-2-hydroxy-1-cyclooctanecarboxamide (8T*) were subjected to X-ray crystal structure analysis. In 8T and 8T*, the hydrogen bonds form infinite ribbons of dimers joined by
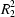
(12) rings with
Ci symmetry. Two types of dimer alternate along each ribbon. The dimers differ in the donor and acceptor roles of the functional groups. This pattern was previously deduced topologically among the possible forms of association for heterochiral dimers [Kálmán
et al. (2002).
Acta Cryst. B
58, 494-501]. As they have the same pattern of hydrogen bonds, 8T and 8T* are isostructural. The additional donor (
i.e. the second hydrogen of the NH
2 group) present in 8T* links the adjacent ribbons so as to form smaller
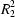
(8) rings between them. The crystals of the
cis stereoisomer 8C are built up from antiparallel hydrogen-bonded helices. The topology and symmetry of this structure are the same as for the close packing of (1
R*,2
R*,4
S*)-4-
tert-butyl-2-hydroxy-1-cyclopentanecarboxamide [Kálmán
et al. (2001).
Acta Cryst. B
57, 539-550]; only the hydrogen-bond donors and acceptors are interchanged, in the same way as in the two dimer types of 8T and 8T* ribbons. This analogy suggests that helices may originate as homochiral dimers with
C2 symmetry and polymerize into helices during crystal formation. The conformational characteristics of the heterochiral dimers observed in the title compounds and in closely related structures are discussed.
Supporting information
CCDC references: 195806; 195807; 195808
For all compounds, data collection: CAD-4 EXPRESS; cell refinement: CAD-4 EXPRESS; data reduction: XCAD4 (Harms 1996); program(s) used to solve structure: SHELXS97 (Sheldrick, 1997); program(s) used to refine structure: SHELXL97 (Sheldrick, 1997).
(8T) (1
R*,2
R*)-
trans-2-hydroxy-1-cyclooctanecarboxylic acid
top
Crystal data top
| C9H16O3 | Z = 2 |
| Mr = 172.22 | F(000) = 188 |
| Triclinic, P1 | Dx = 1.268 Mg m−3 |
| a = 6.035 (1) Å | Mo Kα radiation, λ = 0.710730 Å |
| b = 8.390 (1) Å | Cell parameters from 25 reflections |
| c = 9.389 (2) Å | θ = 17.0–18.9° |
| α = 84.29 (1)° | µ = 0.09 mm−1 |
| β = 76.37 (1)° | T = 293 K |
| γ = 77.95 (1)° | Prism, colourless |
| V = 451.20 (13) Å3 | 0.60 × 0.40 × 0.30 mm |
Data collection top
Enraf-Nonius CAD4
diffractometer | 2585 reflections with I > 2σ(I) |
| Radiation source: fine-focus sealed tube | Rint = 0.021 |
| Graphite monochromator | θmax = 35.0°, θmin = 2.5° |
| ω–2θ scans | h = −9→9 |
Absorption correction: ψ scan
? | k = −13→13 |
| Tmin = 0.935, Tmax = 0.980 | l = −15→15 |
| 8683 measured reflections | 3 standard reflections every 60 min |
| 3931 independent reflections | intensity decay: 3% |
Refinement top
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.044 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.142 | Riding |
| S = 0.95 | w = 1/[σ2(Fo2) + (0.1P)2]
where P = (Fo2 + 2Fc2)/3 |
| 3931 reflections | (Δ/σ)max < 0.001 |
| 111 parameters | Δρmax = 0.33 e Å−3 |
| 101 restraints | Δρmin = −0.20 e Å−3 |
Crystal data top
| C9H16O3 | γ = 77.95 (1)° |
| Mr = 172.22 | V = 451.20 (13) Å3 |
| Triclinic, P1 | Z = 2 |
| a = 6.035 (1) Å | Mo Kα radiation |
| b = 8.390 (1) Å | µ = 0.09 mm−1 |
| c = 9.389 (2) Å | T = 293 K |
| α = 84.29 (1)° | 0.60 × 0.40 × 0.30 mm |
| β = 76.37 (1)° | |
Data collection top
Enraf-Nonius CAD4
diffractometer | 2585 reflections with I > 2σ(I) |
Absorption correction: ψ scan
? | Rint = 0.021 |
| Tmin = 0.935, Tmax = 0.980 | 3 standard reflections every 60 min |
| 8683 measured reflections | intensity decay: 3% |
| 3931 independent reflections | |
Refinement top
| R[F2 > 2σ(F2)] = 0.044 | 101 restraints |
| wR(F2) = 0.142 | Riding |
| S = 0.95 | Δρmax = 0.33 e Å−3 |
| 3931 reflections | Δρmin = −0.20 e Å−3 |
| 111 parameters | |
Special details top
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes)
are estimated using the full covariance matrix. The cell e.s.d.'s are taken
into account individually in the estimation of e.s.d.'s in distances, angles
and torsion angles; correlations between e.s.d.'s in cell parameters are only
used when they are defined by crystal symmetry. An approximate (isotropic)
treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s.
planes. |
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor
wR and goodness of fit S are based on F2, conventional
R-factors R are based on F, with F set to zero for
negative F2. The threshold expression of F2 >
σ(F2) is used only for calculating R-factors(gt) etc.
and is not relevant to the choice of reflections for refinement.
R-factors based on F2 are statistically about twice as large
as those based on F, and R- factors based on ALL data will be
even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| O1 | 0.24663 (10) | 0.42030 (7) | 0.17893 (7) | 0.03759 (15) | |
| H1 | 0.3888 | 0.3949 | 0.1606 | 0.049* | |
| O2 | 0.27899 (11) | 0.66197 (10) | −0.12937 (7) | 0.05039 (19) | |
| O3 | −0.07127 (10) | 0.68921 (8) | 0.01809 (6) | 0.04129 (16) | |
| H3 | −0.1044 | 0.6574 | −0.0526 | 0.054* | |
| C1 | 0.23780 (13) | 0.71043 (9) | 0.12595 (8) | 0.03272 (15) | |
| H1A | 0.4072 | 0.6990 | 0.0995 | 0.043* | |
| C2 | 0.17562 (13) | 0.57618 (9) | 0.24366 (8) | 0.03178 (15) | |
| H2 | 0.0059 | 0.5961 | 0.2756 | 0.041* | |
| C3 | 0.27376 (17) | 0.56596 (10) | 0.38111 (9) | 0.04166 (19) | |
| H3A | 0.4407 | 0.5577 | 0.3504 | 0.054* | |
| H3B | 0.2459 | 0.4659 | 0.4374 | 0.054* | |
| C4 | 0.1765 (2) | 0.70824 (12) | 0.48309 (9) | 0.0492 (2) | |
| H4A | 0.0278 | 0.7635 | 0.4635 | 0.064* | |
| H4B | 0.1480 | 0.6636 | 0.5836 | 0.064* | |
| C5 | 0.3300 (2) | 0.83474 (13) | 0.47017 (11) | 0.0539 (3) | |
| H5A | 0.2344 | 0.9321 | 0.5159 | 0.070* | |
| H5B | 0.4469 | 0.7924 | 0.5267 | 0.070* | |
| C6 | 0.45338 (19) | 0.88506 (16) | 0.31697 (13) | 0.0588 (3) | |
| H6A | 0.5449 | 0.7875 | 0.2696 | 0.076* | |
| H6B | 0.5606 | 0.9529 | 0.3262 | 0.076* | |
| C7 | 0.2998 (3) | 0.97648 (14) | 0.21741 (12) | 0.0593 (3) | |
| H7A | 0.3994 | 1.0105 | 0.1274 | 0.077* | |
| H7B | 0.2102 | 1.0747 | 0.2645 | 0.077* | |
| C8 | 0.1307 (2) | 0.88565 (10) | 0.17615 (10) | 0.0456 (2) | |
| H8A | 0.0667 | 0.9487 | 0.0978 | 0.059* | |
| H8B | 0.0030 | 0.8799 | 0.2603 | 0.059* | |
| C9 | 0.15477 (13) | 0.68416 (9) | −0.00872 (8) | 0.03344 (16) | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| O1 | 0.0362 (3) | 0.0354 (3) | 0.0461 (3) | −0.0020 (2) | −0.0179 (2) | −0.0150 (2) |
| O2 | 0.0391 (3) | 0.0799 (5) | 0.0333 (3) | −0.0071 (3) | −0.0095 (2) | −0.0140 (3) |
| O3 | 0.0323 (3) | 0.0551 (4) | 0.0409 (3) | −0.0039 (2) | −0.0146 (2) | −0.0172 (3) |
| C1 | 0.0320 (3) | 0.0380 (3) | 0.0320 (3) | −0.0069 (3) | −0.0114 (3) | −0.0094 (2) |
| C2 | 0.0305 (3) | 0.0339 (3) | 0.0335 (3) | −0.0029 (2) | −0.0115 (2) | −0.0102 (2) |
| C3 | 0.0531 (5) | 0.0400 (4) | 0.0366 (4) | −0.0029 (3) | −0.0217 (3) | −0.0086 (3) |
| C4 | 0.0655 (6) | 0.0506 (5) | 0.0332 (4) | −0.0059 (4) | −0.0144 (4) | −0.0135 (3) |
| C5 | 0.0657 (6) | 0.0526 (5) | 0.0529 (5) | −0.0023 (4) | −0.0312 (5) | −0.0231 (4) |
| C6 | 0.0445 (5) | 0.0724 (6) | 0.0692 (6) | −0.0208 (5) | −0.0102 (4) | −0.0356 (5) |
| C7 | 0.0882 (8) | 0.0534 (5) | 0.0463 (5) | −0.0365 (6) | −0.0110 (5) | −0.0111 (4) |
| C8 | 0.0629 (6) | 0.0344 (4) | 0.0454 (4) | −0.0056 (4) | −0.0243 (4) | −0.0079 (3) |
| C9 | 0.0329 (3) | 0.0358 (3) | 0.0342 (3) | −0.0029 (3) | −0.0129 (3) | −0.0084 (3) |
Geometric parameters (Å, º) top
| O1—C2 | 1.4369 (8) | C4—C5 | 1.5269 (15) |
| O1—H1 | 0.8200 | C4—H4A | 0.9700 |
| O2—C9 | 1.2096 (10) | C4—H4B | 0.9700 |
| O3—C9 | 1.3199 (10) | C5—C6 | 1.5190 (17) |
| O3—H3 | 0.8200 | C5—H5A | 0.9700 |
| C1—C9 | 1.5161 (10) | C5—H5B | 0.9700 |
| C1—C2 | 1.5376 (12) | C6—C7 | 1.513 (2) |
| C1—C8 | 1.5488 (12) | C6—H6A | 0.9700 |
| C1—H1A | 0.9800 | C6—H6B | 0.9700 |
| C2—C3 | 1.5299 (10) | C7—C8 | 1.5306 (15) |
| C2—H2 | 0.9800 | C7—H7A | 0.9700 |
| C3—C4 | 1.5351 (11) | C7—H7B | 0.9700 |
| C3—H3A | 0.9700 | C8—H8A | 0.9700 |
| C3—H3B | 0.9700 | C8—H8B | 0.9700 |
| | | |
| C2—O1—H1 | 109.5 | C6—C5—H5A | 108.0 |
| C9—O3—H3 | 109.5 | C4—C5—H5A | 108.0 |
| C9—C1—C2 | 108.18 (6) | C6—C5—H5B | 108.0 |
| C9—C1—C8 | 109.36 (6) | C4—C5—H5B | 108.0 |
| C2—C1—C8 | 113.69 (7) | H5A—C5—H5B | 107.2 |
| C9—C1—H1A | 108.5 | C5—C6—C7 | 116.26 (9) |
| C2—C1—H1A | 108.5 | C5—C6—H6A | 108.2 |
| C8—C1—H1A | 108.5 | C7—C6—H6A | 108.2 |
| O1—C2—C1 | 109.35 (6) | C5—C6—H6B | 108.2 |
| O1—C2—C3 | 108.70 (6) | C7—C6—H6B | 108.2 |
| C1—C2—C3 | 116.62 (7) | H6A—C6—H6B | 107.4 |
| O1—C2—H2 | 107.3 | C6—C7—C8 | 117.19 (9) |
| C1—C2—H2 | 107.3 | C6—C7—H7A | 108.0 |
| C3—C2—H2 | 107.3 | C8—C7—H7A | 108.0 |
| C2—C3—C4 | 116.39 (7) | C6—C7—H7B | 108.0 |
| C2—C3—H3A | 108.2 | C8—C7—H7B | 108.0 |
| C4—C3—H3A | 108.2 | H7A—C7—H7B | 107.2 |
| C2—C3—H3B | 108.2 | C7—C8—C1 | 114.96 (9) |
| C4—C3—H3B | 108.2 | C7—C8—H8A | 108.5 |
| H3A—C3—H3B | 107.3 | C1—C8—H8A | 108.5 |
| C5—C4—C3 | 115.80 (9) | C7—C8—H8B | 108.5 |
| C5—C4—H4A | 108.3 | C1—C8—H8B | 108.5 |
| C3—C4—H4A | 108.3 | H8A—C8—H8B | 107.5 |
| C5—C4—H4B | 108.3 | O2—C9—O3 | 122.46 (7) |
| C3—C4—H4B | 108.3 | O2—C9—C1 | 124.59 (7) |
| H4A—C4—H4B | 107.4 | O3—C9—C1 | 112.94 (6) |
| C6—C5—C4 | 117.39 (7) | | |
| | | |
| C9—C1—C2—O1 | 48.02 (8) | C5—C6—C7—C8 | −63.33 (13) |
| C8—C1—C2—O1 | 169.70 (6) | C6—C7—C8—C1 | −47.01 (13) |
| C9—C1—C2—C3 | 171.82 (6) | C9—C1—C8—C7 | −135.91 (8) |
| C8—C1—C2—C3 | −66.51 (9) | C2—C1—C8—C7 | 103.08 (9) |
| O1—C2—C3—C4 | −167.79 (8) | C2—C1—C9—O2 | −121.22 (9) |
| C1—C2—C3—C4 | 68.09 (10) | C8—C1—C9—O2 | 114.48 (10) |
| C2—C3—C4—C5 | −101.20 (10) | C2—C1—C9—O3 | 59.34 (8) |
| C3—C4—C5—C6 | 40.99 (13) | C8—C1—C9—O3 | −64.96 (9) |
| C4—C5—C6—C7 | 66.15 (12) | | |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| O1—H1···O2i | 0.82 | 1.92 | 2.7386 (10) | 177 |
| O3—H3···O1ii | 0.82 | 1.85 | 2.6557 (8) | 166 |
| Symmetry codes: (i) −x+1, −y+1, −z; (ii) −x, −y+1, −z. |
(8Tstar) (1
R*,2
R*)-
trans-2-hydroxy-1-cyclooctanecarboxamide ?
top
Crystal data top
| C9H17NO2 | Z = 2 |
| Mr = 171.24 | F(000) = 188 |
| Triclinic, P1 | Dx = 1.144 Mg m−3 |
| a = 6.760 (1) Å | Mo Kα radiation, λ = 0.710730 Å |
| b = 7.314 (1) Å | Cell parameters from 25 reflections |
| c = 11.217 (1) Å | θ = 12.0–14.4° |
| α = 79.22 (1)° | µ = 0.08 mm−1 |
| β = 74.12 (1)° | T = 293 K |
| γ = 69.45 (1)° | Prism, colourless |
| V = 496.93 (11) Å3 | 0.30 × 0.25 × 0.02 mm |
Data collection top
Enraf-Nonius CAD4
diffractometer | 989 reflections with I > 2σ(I) |
| Radiation source: fine-focus sealed tube | Rint = 0.026 |
| Graphite monochromator | θmax = 26.0°, θmin = 3.0° |
| ω–2θ scans | h = −8→8 |
Absorption correction: ψ scan
? | k = −9→9 |
| Tmin = 0.893, Tmax = 0.995 | l = −13→13 |
| 4315 measured reflections | 3 standard reflections every 60 min |
| 1941 independent reflections | intensity decay: 2% |
Refinement top
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.043 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.146 | Riding |
| S = 0.84 | w = 1/[σ2(Fo2) + (0.1P)2]
where P = (Fo2 + 2Fc2)/3 |
| 1941 reflections | (Δ/σ)max < 0.001 |
| 110 parameters | Δρmax = 0.16 e Å−3 |
| 146 restraints | Δρmin = −0.14 e Å−3 |
Crystal data top
| C9H17NO2 | γ = 69.45 (1)° |
| Mr = 171.24 | V = 496.93 (11) Å3 |
| Triclinic, P1 | Z = 2 |
| a = 6.760 (1) Å | Mo Kα radiation |
| b = 7.314 (1) Å | µ = 0.08 mm−1 |
| c = 11.217 (1) Å | T = 293 K |
| α = 79.22 (1)° | 0.30 × 0.25 × 0.02 mm |
| β = 74.12 (1)° | |
Data collection top
Enraf-Nonius CAD4
diffractometer | 989 reflections with I > 2σ(I) |
Absorption correction: ψ scan
? | Rint = 0.026 |
| Tmin = 0.893, Tmax = 0.995 | 3 standard reflections every 60 min |
| 4315 measured reflections | intensity decay: 2% |
| 1941 independent reflections | |
Refinement top
| R[F2 > 2σ(F2)] = 0.043 | 146 restraints |
| wR(F2) = 0.146 | Riding |
| S = 0.84 | Δρmax = 0.16 e Å−3 |
| 1941 reflections | Δρmin = −0.14 e Å−3 |
| 110 parameters | |
Special details top
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes)
are estimated using the full covariance matrix. The cell e.s.d.'s are taken
into account individually in the estimation of e.s.d.'s in distances, angles
and torsion angles; correlations between e.s.d.'s in cell parameters are only
used when they are defined by crystal symmetry. An approximate (isotropic)
treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s.
planes. |
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor
wR and goodness of fit S are based on F2, conventional
R-factors R are based on F, with F set to zero for
negative F2. The threshold expression of F2 >
σ(F2) is used only for calculating R-factors(gt) etc.
and is not relevant to the choice of reflections for refinement.
R-factors based on F2 are statistically about twice as large
as those based on F, and R- factors based on ALL data will be
even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| O1 | 0.7699 (2) | 0.3358 (2) | 0.52719 (13) | 0.0553 (4) | |
| H1 | 0.6963 | 0.3068 | 0.4911 | 0.072* | |
| O2 | 0.4304 (2) | 0.79742 (19) | 0.60322 (14) | 0.0552 (4) | |
| N1 | 0.7632 (3) | 0.8183 (2) | 0.52684 (17) | 0.0550 (5) | |
| H1B | 0.7152 | 0.9264 | 0.4825 | 0.071* | |
| H1C | 0.8997 | 0.7686 | 0.5245 | 0.071* | |
| C1 | 0.7208 (3) | 0.5425 (3) | 0.67721 (19) | 0.0455 (5) | |
| H1A | 0.8794 | 0.5032 | 0.6519 | 0.059* | |
| C2 | 0.6472 (3) | 0.3823 (3) | 0.65028 (19) | 0.0497 (5) | |
| H2 | 0.4946 | 0.4419 | 0.6459 | 0.065* | |
| C3 | 0.6631 (4) | 0.2002 (3) | 0.7427 (2) | 0.0630 (6) | |
| H3A | 0.5780 | 0.2422 | 0.8235 | 0.082* | |
| H3B | 0.5941 | 0.1215 | 0.7183 | 0.082* | |
| C4 | 0.8849 (4) | 0.0676 (3) | 0.7595 (3) | 0.0708 (7) | |
| H4A | 0.9677 | 0.0190 | 0.6800 | 0.092* | |
| H4B | 0.8660 | −0.0447 | 0.8172 | 0.092* | |
| C5 | 1.0213 (4) | 0.1550 (4) | 0.8062 (3) | 0.0758 (8) | |
| H5A | 1.1439 | 0.0489 | 0.8286 | 0.099* | |
| H5B | 1.0784 | 0.2394 | 0.7383 | 0.099* | |
| C6 | 0.9013 (5) | 0.2732 (4) | 0.9180 (3) | 0.0872 (9) | |
| H6A | 0.9957 | 0.2398 | 0.9756 | 0.113* | |
| H6B | 0.7754 | 0.2327 | 0.9604 | 0.113* | |
| C7 | 0.8258 (5) | 0.4964 (4) | 0.8882 (2) | 0.0793 (8) | |
| H7A | 0.7742 | 0.5544 | 0.9665 | 0.103* | |
| H7B | 0.9518 | 0.5347 | 0.8425 | 0.103* | |
| C8 | 0.6510 (4) | 0.5874 (3) | 0.8147 (2) | 0.0638 (6) | |
| H8A | 0.6021 | 0.7286 | 0.8166 | 0.083* | |
| H8B | 0.5285 | 0.5415 | 0.8559 | 0.083* | |
| C9 | 0.6286 (3) | 0.7294 (3) | 0.59782 (18) | 0.0436 (5) | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| O1 | 0.0512 (8) | 0.0545 (10) | 0.0644 (10) | −0.0112 (7) | −0.0224 (7) | −0.0130 (7) |
| O2 | 0.0439 (8) | 0.0441 (8) | 0.0742 (10) | −0.0057 (6) | −0.0221 (7) | −0.0010 (7) |
| N1 | 0.0496 (10) | 0.0415 (10) | 0.0670 (12) | −0.0095 (8) | −0.0187 (9) | 0.0093 (9) |
| C1 | 0.0469 (11) | 0.0388 (11) | 0.0511 (12) | −0.0100 (9) | −0.0177 (9) | −0.0023 (9) |
| C2 | 0.0432 (10) | 0.0438 (11) | 0.0594 (13) | −0.0078 (9) | −0.0167 (10) | −0.0033 (10) |
| C3 | 0.0656 (14) | 0.0465 (13) | 0.0777 (16) | −0.0201 (11) | −0.0193 (12) | 0.0007 (11) |
| C4 | 0.0827 (16) | 0.0475 (13) | 0.0781 (17) | −0.0155 (12) | −0.0291 (14) | 0.0095 (12) |
| C5 | 0.0811 (17) | 0.0616 (16) | 0.0823 (18) | −0.0118 (13) | −0.0406 (15) | 0.0099 (14) |
| C6 | 0.117 (2) | 0.0747 (19) | 0.0737 (18) | −0.0218 (17) | −0.0511 (16) | 0.0096 (15) |
| C7 | 0.112 (2) | 0.0728 (18) | 0.0606 (16) | −0.0252 (16) | −0.0388 (15) | −0.0034 (13) |
| C8 | 0.0838 (16) | 0.0481 (13) | 0.0564 (15) | −0.0152 (12) | −0.0179 (12) | −0.0059 (11) |
| C9 | 0.0489 (11) | 0.0357 (10) | 0.0473 (12) | −0.0070 (9) | −0.0183 (9) | −0.0081 (9) |
Geometric parameters (Å, º) top
| O1—C2 | 1.436 (2) | C4—C5 | 1.529 (4) |
| O1—H1 | 0.8200 | C4—H4A | 0.9700 |
| O2—C9 | 1.243 (2) | C4—H4B | 0.9700 |
| N1—C9 | 1.307 (3) | C5—C6 | 1.527 (4) |
| N1—H1B | 0.8600 | C5—H5A | 0.9700 |
| N1—H1C | 0.8600 | C5—H5B | 0.9700 |
| C1—C9 | 1.517 (3) | C6—C7 | 1.530 (4) |
| C1—C2 | 1.528 (3) | C6—H6A | 0.9700 |
| C1—C8 | 1.546 (3) | C6—H6B | 0.9700 |
| C1—H1A | 0.9800 | C7—C8 | 1.515 (3) |
| C2—C3 | 1.518 (3) | C7—H7A | 0.9700 |
| C2—H2 | 0.9800 | C7—H7B | 0.9700 |
| C3—C4 | 1.513 (3) | C8—H8A | 0.9700 |
| C3—H3A | 0.9700 | C8—H8B | 0.9700 |
| C3—H3B | 0.9700 | | |
| | | |
| C2—O1—H1 | 109.5 | C6—C5—C4 | 115.2 (2) |
| C9—N1—H1B | 120.0 | C6—C5—H5A | 108.5 |
| C9—N1—H1C | 120.0 | C4—C5—H5A | 108.5 |
| H1B—N1—H1C | 120.0 | C6—C5—H5B | 108.5 |
| C9—C1—C2 | 107.95 (16) | C4—C5—H5B | 108.5 |
| C9—C1—C8 | 108.31 (16) | H5A—C5—H5B | 107.5 |
| C2—C1—C8 | 115.21 (18) | C5—C6—C7 | 115.7 (2) |
| C9—C1—H1A | 108.4 | C5—C6—H6A | 108.4 |
| C2—C1—H1A | 108.4 | C7—C6—H6A | 108.4 |
| C8—C1—H1A | 108.4 | C5—C6—H6B | 108.4 |
| O1—C2—C3 | 112.02 (18) | C7—C6—H6B | 108.4 |
| O1—C2—C1 | 104.85 (16) | H6A—C6—H6B | 107.4 |
| C3—C2—C1 | 117.76 (18) | C8—C7—C6 | 118.3 (2) |
| O1—C2—H2 | 107.2 | C8—C7—H7A | 107.7 |
| C3—C2—H2 | 107.2 | C6—C7—H7A | 107.7 |
| C1—C2—H2 | 107.2 | C8—C7—H7B | 107.7 |
| C4—C3—C2 | 118.8 (2) | C6—C7—H7B | 107.7 |
| C4—C3—H3A | 107.6 | H7A—C7—H7B | 107.1 |
| C2—C3—H3A | 107.6 | C7—C8—C1 | 115.1 (2) |
| C4—C3—H3B | 107.6 | C7—C8—H8A | 108.5 |
| C2—C3—H3B | 107.6 | C1—C8—H8A | 108.5 |
| H3A—C3—H3B | 107.1 | C7—C8—H8B | 108.5 |
| C3—C4—C5 | 117.7 (2) | C1—C8—H8B | 108.5 |
| C3—C4—H4A | 107.9 | H8A—C8—H8B | 107.5 |
| C5—C4—H4A | 107.9 | O2—C9—N1 | 121.56 (17) |
| C3—C4—H4B | 107.9 | O2—C9—C1 | 120.54 (18) |
| C5—C4—H4B | 107.9 | N1—C9—C1 | 117.87 (16) |
| H4A—C4—H4B | 107.2 | | |
| | | |
| C9—C1—C2—O1 | 71.69 (18) | C5—C6—C7—C8 | 66.8 (4) |
| C8—C1—C2—O1 | −167.14 (16) | C6—C7—C8—C1 | −68.7 (3) |
| C9—C1—C2—C3 | −163.01 (18) | C9—C1—C8—C7 | −138.0 (2) |
| C8—C1—C2—C3 | −41.8 (3) | C2—C1—C8—C7 | 101.0 (2) |
| O1—C2—C3—C4 | 57.1 (3) | C2—C1—C9—O2 | 55.2 (2) |
| C1—C2—C3—C4 | −64.6 (3) | C8—C1—C9—O2 | −70.2 (2) |
| C2—C3—C4—C5 | 61.3 (3) | C2—C1—C9—N1 | −126.63 (19) |
| C3—C4—C5—C6 | 46.7 (3) | C8—C1—C9—N1 | 108.0 (2) |
| C4—C5—C6—C7 | −100.3 (3) | | |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| O1—H1···O2i | 0.82 | 1.92 | 2.732 (2) | 170 |
| N1—H1B···O2ii | 0.86 | 2.09 | 2.945 (2) | 172 |
| N1—H1C···O1iii | 0.86 | 2.04 | 2.883 (2) | 166 |
| Symmetry codes: (i) −x+1, −y+1, −z+1; (ii) −x+1, −y+2, −z+1; (iii) −x+2, −y+1, −z+1. |
(8C) (1
R*,2
S*)-
cis-2-hydroxy-1-cyclooctanecarboxylic acid
top
Crystal data top
| C9H16O3 | F(000) = 376 |
| Mr = 172.22 | Dx = 1.215 Mg m−3 |
| Monoclinic, P21/c | Mo Kα radiation, λ = 0.710730 Å |
| a = 11.082 (1) Å | Cell parameters from 25 reflections |
| b = 7.618 (1) Å | θ = 16.2–17.8° |
| c = 11.579 (1) Å | µ = 0.09 mm−1 |
| β = 105.67 (1)° | T = 293 K |
| V = 941.20 (17) Å3 | Prism, colourless |
| Z = 4 | 0.50 × 0.40 × 0.25 mm |
Data collection top
Enraf-Nonius CAD4
diffractometer | 2100 reflections with I > 2σ(I) |
| Radiation source: fine-focus sealed tube | Rint = 0.016 |
| Graphite monochromator | θmax = 35.0°, θmin = 3.2° |
| ω–2θ scans | h = −17→17 |
Absorption correction: ψ scan
? | k = −12→0 |
| Tmin = 0.958, Tmax = 0.988 | l = 0→18 |
| 4372 measured reflections | 3 standard reflections every 60 min |
| 4074 independent reflections | intensity decay: 3% |
Refinement top
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.050 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.154 | Riding |
| S = 0.85 | w = 1/[σ2(Fo2) + (0.1P)2]
where P = (Fo2 + 2Fc2)/3 |
| 4074 reflections | (Δ/σ)max < 0.001 |
| 111 parameters | Δρmax = 0.31 e Å−3 |
| 99 restraints | Δρmin = −0.17 e Å−3 |
Crystal data top
| C9H16O3 | V = 941.20 (17) Å3 |
| Mr = 172.22 | Z = 4 |
| Monoclinic, P21/c | Mo Kα radiation |
| a = 11.082 (1) Å | µ = 0.09 mm−1 |
| b = 7.618 (1) Å | T = 293 K |
| c = 11.579 (1) Å | 0.50 × 0.40 × 0.25 mm |
| β = 105.67 (1)° | |
Data collection top
Enraf-Nonius CAD4
diffractometer | 2100 reflections with I > 2σ(I) |
Absorption correction: ψ scan
? | Rint = 0.016 |
| Tmin = 0.958, Tmax = 0.988 | 3 standard reflections every 60 min |
| 4372 measured reflections | intensity decay: 3% |
| 4074 independent reflections | |
Refinement top
| R[F2 > 2σ(F2)] = 0.050 | 99 restraints |
| wR(F2) = 0.154 | Riding |
| S = 0.85 | Δρmax = 0.31 e Å−3 |
| 4074 reflections | Δρmin = −0.17 e Å−3 |
| 111 parameters | |
Special details top
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes)
are estimated using the full covariance matrix. The cell e.s.d.'s are taken
into account individually in the estimation of e.s.d.'s in distances, angles
and torsion angles; correlations between e.s.d.'s in cell parameters are only
used when they are defined by crystal symmetry. An approximate (isotropic)
treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s.
planes. |
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor
wR and goodness of fit S are based on F2, conventional
R-factors R are based on F, with F set to zero for
negative F2. The threshold expression of F2 >
σ(F2) is used only for calculating R-factors(gt) etc.
and is not relevant to the choice of reflections for refinement.
R-factors based on F2 are statistically about twice as large
as those based on F, and R- factors based on ALL data will be
even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| O1 | 0.00245 (6) | 0.22080 (10) | 0.11454 (7) | 0.03749 (19) | |
| H1 | −0.0107 | 0.1985 | 0.0429 | 0.049* | |
| O2 | 0.07036 (8) | −0.17111 (10) | 0.13317 (7) | 0.0418 (2) | |
| O3 | 0.14942 (8) | −0.06238 (11) | 0.31595 (7) | 0.0475 (2) | |
| H3 | 0.1048 | −0.1382 | 0.3333 | 0.062* | |
| C1 | 0.20784 (9) | 0.08228 (13) | 0.15978 (8) | 0.03058 (19) | |
| H1A | 0.2892 | 0.0920 | 0.2195 | 0.040* | |
| C2 | 0.13409 (10) | 0.25314 (13) | 0.16540 (9) | 0.0339 (2) | |
| H2 | 0.1465 | 0.2825 | 0.2502 | 0.044* | |
| C3 | 0.17177 (13) | 0.41215 (15) | 0.10455 (13) | 0.0490 (3) | |
| H3A | 0.1116 | 0.5047 | 0.1042 | 0.064* | |
| H3B | 0.1643 | 0.3816 | 0.0216 | 0.064* | |
| C4 | 0.30256 (15) | 0.4869 (2) | 0.15783 (16) | 0.0659 (4) | |
| H4A | 0.3075 | 0.5981 | 0.1186 | 0.086* | |
| H4B | 0.3111 | 0.5121 | 0.2418 | 0.086* | |
| C5 | 0.41537 (15) | 0.3764 (3) | 0.15052 (16) | 0.0709 (4) | |
| H5A | 0.4314 | 0.2906 | 0.2148 | 0.092* | |
| H5B | 0.4880 | 0.4528 | 0.1653 | 0.092* | |
| C6 | 0.40454 (14) | 0.2802 (2) | 0.03419 (15) | 0.0679 (4) | |
| H6A | 0.3448 | 0.3425 | −0.0293 | 0.088* | |
| H6B | 0.4851 | 0.2857 | 0.0165 | 0.088* | |
| C7 | 0.36447 (12) | 0.0883 (2) | 0.03002 (13) | 0.0558 (3) | |
| H7A | 0.4235 | 0.0279 | 0.0951 | 0.073* | |
| H7B | 0.3739 | 0.0387 | −0.0442 | 0.073* | |
| C8 | 0.23276 (10) | 0.04339 (15) | 0.03808 (10) | 0.0389 (2) | |
| H8A | 0.2178 | −0.0805 | 0.0205 | 0.051* | |
| H8B | 0.1728 | 0.1087 | −0.0234 | 0.051* | |
| C9 | 0.13589 (9) | −0.06453 (13) | 0.19924 (9) | 0.0317 (2) | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| O1 | 0.0377 (4) | 0.0430 (4) | 0.0329 (4) | 0.0057 (3) | 0.0116 (3) | −0.0072 (3) |
| O2 | 0.0545 (5) | 0.0348 (4) | 0.0360 (4) | −0.0087 (3) | 0.0123 (3) | −0.0054 (3) |
| O3 | 0.0591 (5) | 0.0547 (5) | 0.0298 (4) | −0.0231 (4) | 0.0142 (3) | −0.0013 (3) |
| C1 | 0.0319 (4) | 0.0335 (4) | 0.0272 (4) | −0.0001 (4) | 0.0094 (3) | 0.0011 (3) |
| C2 | 0.0404 (5) | 0.0325 (5) | 0.0282 (4) | 0.0006 (4) | 0.0084 (4) | −0.0035 (4) |
| C3 | 0.0580 (7) | 0.0342 (5) | 0.0534 (7) | −0.0001 (5) | 0.0123 (6) | 0.0077 (5) |
| C4 | 0.0755 (10) | 0.0462 (7) | 0.0744 (10) | −0.0216 (7) | 0.0175 (8) | −0.0033 (7) |
| C5 | 0.0572 (8) | 0.0814 (10) | 0.0695 (10) | −0.0275 (8) | 0.0091 (7) | 0.0028 (8) |
| C6 | 0.0537 (8) | 0.0890 (11) | 0.0685 (9) | −0.0161 (8) | 0.0292 (7) | 0.0076 (8) |
| C7 | 0.0439 (6) | 0.0772 (9) | 0.0537 (7) | 0.0001 (6) | 0.0258 (6) | −0.0004 (7) |
| C8 | 0.0401 (5) | 0.0469 (6) | 0.0341 (5) | 0.0010 (4) | 0.0177 (4) | −0.0025 (4) |
| C9 | 0.0353 (4) | 0.0314 (4) | 0.0291 (4) | 0.0016 (4) | 0.0100 (3) | 0.0007 (3) |
Geometric parameters (Å, º) top
| O1—C2 | 1.4393 (12) | C4—C5 | 1.528 (3) |
| O1—H1 | 0.8200 | C4—H4A | 0.9700 |
| O2—C9 | 1.2131 (12) | C4—H4B | 0.9700 |
| O3—C9 | 1.3189 (12) | C5—C6 | 1.509 (2) |
| O3—H3 | 0.8200 | C5—H5A | 0.9700 |
| C1—C9 | 1.5140 (14) | C5—H5B | 0.9700 |
| C1—C8 | 1.5363 (13) | C6—C7 | 1.525 (2) |
| C1—C2 | 1.5476 (14) | C6—H6A | 0.9700 |
| C1—H1A | 0.9800 | C6—H6B | 0.9700 |
| C2—C3 | 1.5157 (16) | C7—C8 | 1.5263 (16) |
| C2—H2 | 0.9800 | C7—H7A | 0.9700 |
| C3—C4 | 1.5239 (19) | C7—H7B | 0.9700 |
| C3—H3A | 0.9700 | C8—H8A | 0.9700 |
| C3—H3B | 0.9700 | C8—H8B | 0.9700 |
| | | |
| C2—O1—H1 | 109.5 | C4—C5—H5A | 108.1 |
| C9—O3—H3 | 109.5 | C6—C5—H5A | 108.1 |
| C9—C1—C8 | 111.98 (8) | C4—C5—H5B | 108.1 |
| C9—C1—C2 | 106.34 (8) | C6—C5—H5B | 108.1 |
| C8—C1—C2 | 115.38 (8) | H5A—C5—H5B | 107.3 |
| C9—C1—H1A | 107.6 | C7—C6—C5 | 116.50 (13) |
| C8—C1—H1A | 107.6 | C7—C6—H6A | 108.2 |
| C2—C1—H1A | 107.6 | C5—C6—H6A | 108.2 |
| O1—C2—C3 | 108.82 (9) | C7—C6—H6B | 108.2 |
| O1—C2—C1 | 109.10 (8) | C5—C6—H6B | 108.2 |
| C3—C2—C1 | 115.96 (9) | H6A—C6—H6B | 107.3 |
| O1—C2—H2 | 107.5 | C6—C7—C8 | 119.21 (12) |
| C3—C2—H2 | 107.5 | C6—C7—H7A | 107.5 |
| C1—C2—H2 | 107.5 | C8—C7—H7A | 107.5 |
| C2—C3—C4 | 117.49 (11) | C6—C7—H7B | 107.5 |
| C2—C3—H3A | 107.9 | C8—C7—H7B | 107.5 |
| C4—C3—H3A | 107.9 | H7A—C7—H7B | 107.0 |
| C2—C3—H3B | 107.9 | C7—C8—C1 | 115.13 (10) |
| C4—C3—H3B | 107.9 | C7—C8—H8A | 108.5 |
| H3A—C3—H3B | 107.2 | C1—C8—H8A | 108.5 |
| C5—C4—C3 | 118.33 (13) | C7—C8—H8B | 108.5 |
| C5—C4—H4A | 107.7 | C1—C8—H8B | 108.5 |
| C3—C4—H4A | 107.7 | H8A—C8—H8B | 107.5 |
| C5—C4—H4B | 107.7 | O2—C9—O3 | 122.51 (9) |
| C3—C4—H4B | 107.7 | O2—C9—C1 | 125.34 (9) |
| H4A—C4—H4B | 107.1 | O3—C9—C1 | 112.13 (8) |
| C4—C5—C6 | 116.60 (14) | | |
| | | |
| C9—C1—C2—O1 | −43.91 (11) | C5—C6—C7—C8 | 66.10 (19) |
| C8—C1—C2—O1 | 80.90 (11) | C6—C7—C8—C1 | −67.83 (17) |
| C9—C1—C2—C3 | −167.16 (9) | C9—C1—C8—C7 | −137.14 (11) |
| C8—C1—C2—C3 | −42.36 (13) | C2—C1—C8—C7 | 101.04 (12) |
| O1—C2—C3—C4 | 170.74 (11) | C8—C1—C9—O2 | −22.54 (14) |
| C1—C2—C3—C4 | −65.86 (14) | C2—C1—C9—O2 | 104.33 (11) |
| C2—C3—C4—C5 | 67.30 (18) | C8—C1—C9—O3 | 159.09 (9) |
| C3—C4—C5—C6 | 41.1 (2) | C2—C1—C9—O3 | −74.04 (10) |
| C4—C5—C6—C7 | −96.78 (17) | | |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| O1—H1···O2i | 0.82 | 1.98 | 2.7876 (11) | 170 |
| O3—H3···O1ii | 0.82 | 1.82 | 2.6344 (11) | 171 |
| Symmetry codes: (i) −x, −y, −z; (ii) −x, y−1/2, −z+1/2. |
Experimental details
| | (8T) | (8Tstar) | (8C) |
| Crystal data |
| Chemical formula | C9H16O3 | C9H17NO2 | C9H16O3 |
| Mr | 172.22 | 171.24 | 172.22 |
| Crystal system, space group | Triclinic, P1 | Triclinic, P1 | Monoclinic, P21/c |
| Temperature (K) | 293 | 293 | 293 |
| a, b, c (Å) | 6.035 (1), 8.390 (1), 9.389 (2) | 6.760 (1), 7.314 (1), 11.217 (1) | 11.082 (1), 7.618 (1), 11.579 (1) |
| α, β, γ (°) | 84.29 (1), 76.37 (1), 77.95 (1) | 79.22 (1), 74.12 (1), 69.45 (1) | 90, 105.67 (1), 90 |
| V (Å3) | 451.20 (13) | 496.93 (11) | 941.20 (17) |
| Z | 2 | 2 | 4 |
| Radiation type | Mo Kα | Mo Kα | Mo Kα |
| µ (mm−1) | 0.09 | 0.08 | 0.09 |
| Crystal size (mm) | 0.60 × 0.40 × 0.30 | 0.30 × 0.25 × 0.02 | 0.50 × 0.40 × 0.25 |
| |
| Data collection |
| Diffractometer | Enraf-Nonius CAD4
diffractometer | Enraf-Nonius CAD4
diffractometer | Enraf-Nonius CAD4
diffractometer |
| Absorption correction | ψ scan | ψ scan | ψ scan |
| Tmin, Tmax | 0.935, 0.980 | 0.893, 0.995 | 0.958, 0.988 |
No. of measured, independent and
observed [I > 2σ(I)] reflections | 8683, 3931, 2585 | 4315, 1941, 989 | 4372, 4074, 2100 |
| Rint | 0.021 | 0.026 | 0.016 |
| (sin θ/λ)max (Å−1) | 0.806 | 0.617 | 0.806 |
| |
| Refinement |
| R[F2 > 2σ(F2)], wR(F2), S | 0.044, 0.142, 0.95 | 0.043, 0.146, 0.84 | 0.050, 0.154, 0.85 |
| No. of reflections | 3931 | 1941 | 4074 |
| No. of parameters | 111 | 110 | 111 |
| No. of restraints | 101 | 146 | 99 |
| H-atom treatment | Riding | Riding | Riding |
| Δρmax, Δρmin (e Å−3) | 0.33, −0.20 | 0.16, −0.14 | 0.31, −0.17 |
Hydrogen-bond geometry (Å, º) for (8T) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| O1—H1···O2i | 0.82 | 1.92 | 2.7386 (10) | 176.6 |
| O3—H3···O1ii | 0.82 | 1.85 | 2.6557 (8) | 165.7 |
| Symmetry codes: (i) −x+1, −y+1, −z; (ii) −x, −y+1, −z. |
Hydrogen-bond geometry (Å, º) for (8Tstar) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| O1—H1···O2i | 0.82 | 1.92 | 2.732 (2) | 170.0 |
| N1—H1B···O2ii | 0.86 | 2.09 | 2.945 (2) | 172.3 |
| N1—H1C···O1iii | 0.86 | 2.04 | 2.883 (2) | 165.9 |
| Symmetry codes: (i) −x+1, −y+1, −z+1; (ii) −x+1, −y+2, −z+1; (iii) −x+2, −y+1, −z+1. |
Hydrogen-bond geometry (Å, º) for (8C) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| O1—H1···O2i | 0.82 | 1.98 | 2.7876 (11) | 169.5 |
| O3—H3···O1ii | 0.82 | 1.82 | 2.6344 (11) | 170.9 |
| Symmetry codes: (i) −x, −y, −z; (ii) −x, y−1/2, −z+1/2. |

 Subscribe to Acta Crystallographica Section B: Structural Science, Crystal Engineering and Materials
Subscribe to Acta Crystallographica Section B: Structural Science, Crystal Engineering and Materials
The full text of this article is available to subscribers to the journal.
If you have already registered and are using a computer listed in your registration details, please email
support@iucr.org for assistance.

 Subscribe to Acta Crystallographica Section B: Structural Science, Crystal Engineering and Materials
Subscribe to Acta Crystallographica Section B: Structural Science, Crystal Engineering and Materials
 journal menu
journal menu








![[Figure 1]](http://journals.iucr.org/b/issues/2002/05/00/de0017/de0017fig1thm.gif)
![[Figure 2]](http://journals.iucr.org/b/issues/2002/05/00/de0017/de0017fig2thm.gif)
![[Figure 3]](http://journals.iucr.org/b/issues/2002/05/00/de0017/de0017fig3thm.gif)
![[Figure 4]](http://journals.iucr.org/b/issues/2002/05/00/de0017/de0017fig4thm.gif)
![[Figure 5]](http://journals.iucr.org/b/issues/2002/05/00/de0017/de0017fig5thm.gif)
![[Figure 6]](http://journals.iucr.org/b/issues/2002/05/00/de0017/de0017fig6thm.gif)
![[Figure 7]](http://journals.iucr.org/b/issues/2002/05/00/de0017/de0017fig7thm.gif)
![[Figure 8]](http://journals.iucr.org/b/issues/2002/05/00/de0017/de0017fig8thm.gif)
![[Figure 9]](http://journals.iucr.org/b/issues/2002/05/00/de0017/de0017fig9thm.gif)

 Subscribe to Acta Crystallographica Section B: Structural Science, Crystal Engineering and Materials
Subscribe to Acta Crystallographica Section B: Structural Science, Crystal Engineering and Materials


