Buy article online - an online subscription or single-article purchase is required to access this article.

Two acylhydrazone complexes, bis{6-methyl-
N′-[1-(pyrazin-2-yl-κ
N1)ethylidene]nicotinohydrazidato-κ
2N′,
O}nickel(II), [Ni(C
13H
12N
5O)
2], (I), and di-μ-azido-κ
4N1:
N1-bis({6-methyl-
N′-[1-(pyrazin-2-yl-κ
N1)ethylidene]nicotinohydrazidato-κ
2N′,
O}nickel(II)), [Cu
2(C
13H
12N
5O)
2(N
3)
2], (II), derived from 6-methyl-
N′-[1-(pyrazin-2-yl)ethylidene]nicotinohydrazide (H
L) and azide salts, have been synthesized. H
L acts as an
N,
N′,
O-tridentate ligand in both complexes. Complex (I) crystallizes in the orthorhombic space group
Pbcn and has a mononuclear structure, the azide co-ligand is not involved in crystallization and the Ni
2+ centre lies in a distorted {N
4O
2} octahedral coordination environment. Complex (II) crystallizes in the triclinic space group
P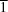
and is a centrosymmetric binuclear complex with a crystallographically independent Cu
2+ centre coordinating to three donor atoms from the deprotonated
L− ligand and to two N atoms belonging to two bridging azide anions. The two- and one-dimensional supramolecular structures are constructed by hydrogen-bonding interactions in (I) and (II), respectively. The
in vitro urease inhibitory evaluation revealed that complex (II) showed a better inhibitory activity, with the IC
50 value being 1.32±0.4 µ
M. Both complexes can effectively bind to bovine serum albumin (BSA) by 1:1 binding, which was assessed
via tryptophan emission–quenching measurements. The bioactivities of the two complexes towards
jack bean urease were also studied by molecular docking. The effects of the metal ions and the coordination environments in the two complexes on
in vitro urease inhibitory activity are preliminarily discussed.
Supporting information
CCDC references: 1920797; 1920796
For both structures, data collection: CrysAlis PRO (Agilent, 2014); cell refinement: CrysAlis PRO (Agilent, 2014); data reduction: CrysAlis PRO (Agilent, 2014); program(s) used to solve structure: SHELXS97 (Sheldrick, 2008); program(s) used to refine structure: SHELXL97 (Sheldrick, 2008); molecular graphics: OLEX2 (Dolomanov et al., 2009); software used to prepare material for publication: OLEX2 (Dolomanov et al., 2009).
Bis{6-methyl-
N'-[1-(pyrazin-2-yl-
κN1)ethylidene]nicotinohydrazidato-
κ2N',
O}nickel(II) (I)
top
Crystal data top
| [Ni(C13H12N5O)2] | Dx = 1.439 Mg m−3 |
| Mr = 567.26 | Mo Kα radiation, λ = 0.71073 Å |
| Orthorhombic, Pbcn | Cell parameters from 8956 reflections |
| a = 11.9385 (6) Å | θ = 2.5–26.2° |
| b = 9.7023 (4) Å | µ = 0.79 mm−1 |
| c = 22.5975 (10) Å | T = 273 K |
| V = 2617.5 (2) Å3 | Block, green |
| Z = 4 | 0.24 × 0.15 × 0.13 mm |
| F(000) = 1176 | |
Data collection top
CCD area detector
diffractometer | 2307 independent reflections |
| Radiation source: fine-focus sealed tube | 1720 reflections with I > 2σ(I) |
| Graphite monochromator | Rint = 0.047 |
| phi and ω scans | θmax = 25.0°, θmin = 2.5° |
Absorption correction: multi-scan
(CrysAlis PRO; Agilent, 2014) | h = −14→14 |
| Tmin = 0.834, Tmax = 0.905 | k = −11→9 |
| 23203 measured reflections | l = −26→26 |
Refinement top
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.045 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.124 | H atoms treated by a mixture of independent and constrained refinement |
| S = 1.05 | w = 1/[σ2(Fo2) + (0.0569P)2 + 2.7448P]
where P = (Fo2 + 2Fc2)/3 |
| 2307 reflections | (Δ/σ)max < 0.001 |
| 179 parameters | Δρmax = 0.40 e Å−3 |
| 0 restraints | Δρmin = −0.18 e Å−3 |
Special details top
Geometry. All esds (except the esd in the dihedral angle between two l.s. planes)
are estimated using the full covariance matrix. The cell esds are taken
into account individually in the estimation of esds in distances, angles
and torsion angles; correlations between esds in cell parameters are only
used when they are defined by crystal symmetry. An approximate (isotropic)
treatment of cell esds is used for estimating esds involving l.s. planes. |
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and
goodness of fit S are based on F2, conventional R-factors R are based
on F, with F set to zero for negative F2. The threshold expression of
F2 > 2sigma(F2) is used only for calculating R-factors(gt) etc. and is
not relevant to the choice of reflections for refinement. R-factors based
on F2 are statistically about twice as large as those based on F, and R-
factors based on ALL data will be even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| Ni1 | 0.0000 | 0.24251 (5) | 0.7500 | 0.0356 (2) | |
| C1 | 0.0214 (3) | 0.4757 (4) | 0.84393 (15) | 0.0533 (9) | |
| H1 | −0.0558 | 0.4708 | 0.8493 | 0.064* | |
| C2 | 0.0821 (4) | 0.5697 (4) | 0.87618 (16) | 0.0664 (10) | |
| H2 | 0.0449 | 0.6257 | 0.9032 | 0.080* | |
| C3 | 0.2404 (3) | 0.4995 (3) | 0.83124 (15) | 0.0529 (8) | |
| H3 | 0.3174 | 0.5062 | 0.8258 | 0.063* | |
| C4 | 0.1818 (2) | 0.4031 (3) | 0.79856 (13) | 0.0402 (7) | |
| C5 | 0.2322 (2) | 0.3108 (3) | 0.75412 (12) | 0.0379 (7) | |
| C6 | 0.3558 (3) | 0.3024 (4) | 0.74455 (16) | 0.0560 (9) | |
| H6A | 0.3894 | 0.2507 | 0.7762 | 0.084* | |
| H6B | 0.3869 | 0.3936 | 0.7438 | 0.084* | |
| H6C | 0.3706 | 0.2573 | 0.7076 | 0.084* | |
| C7 | 0.0996 (3) | 0.0900 (3) | 0.66016 (13) | 0.0434 (7) | |
| C8 | 0.1161 (3) | −0.0018 (3) | 0.60861 (14) | 0.0550 (9) | |
| C9 | 0.2162 (4) | −0.0139 (5) | 0.57894 (19) | 0.0853 (13) | |
| H9 | 0.2780 | 0.0384 | 0.5902 | 0.102* | |
| C10 | 0.2234 (5) | −0.1051 (6) | 0.5322 (2) | 0.1057 (18) | |
| H10 | 0.2910 | −0.1157 | 0.5122 | 0.127* | |
| C11 | 0.1331 (6) | −0.1789 (4) | 0.51517 (19) | 0.1000 (19) | |
| C12 | 0.0303 (5) | −0.0817 (5) | 0.5879 (2) | 0.0993 (17) | |
| H12 | −0.0385 | −0.0749 | 0.6070 | 0.119* | |
| C13 | 0.1392 (7) | −0.2780 (5) | 0.4635 (2) | 0.146 (3) | |
| H13A | 0.0840 | −0.3491 | 0.4684 | 0.220* | |
| H13B | 0.2124 | −0.3187 | 0.4619 | 0.220* | |
| H13C | 0.1250 | −0.2290 | 0.4273 | 0.220* | |
| N1 | 0.06958 (19) | 0.3924 (2) | 0.80568 (10) | 0.0404 (6) | |
| N2 | 0.1916 (3) | 0.5830 (3) | 0.87003 (14) | 0.0675 (9) | |
| N3 | 0.1596 (2) | 0.2387 (2) | 0.72554 (12) | 0.0385 (6) | |
| N4 | 0.1912 (2) | 0.1529 (3) | 0.68041 (11) | 0.0444 (6) | |
| N5 | 0.0378 (5) | −0.1688 (5) | 0.54224 (18) | 0.1206 (18) | |
| O1 | 0.00128 (16) | 0.1004 (2) | 0.68171 (10) | 0.0454 (5) | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| Ni1 | 0.0229 (3) | 0.0408 (3) | 0.0432 (3) | 0.000 | 0.0000 (2) | 0.000 |
| C1 | 0.053 (2) | 0.054 (2) | 0.053 (2) | 0.0030 (16) | 0.0068 (16) | −0.0048 (17) |
| C2 | 0.086 (3) | 0.056 (2) | 0.057 (2) | 0.003 (2) | 0.003 (2) | −0.0080 (18) |
| C3 | 0.0490 (19) | 0.054 (2) | 0.0556 (19) | −0.0142 (17) | −0.0106 (17) | 0.0053 (17) |
| C4 | 0.0344 (16) | 0.0405 (16) | 0.0455 (17) | −0.0057 (13) | −0.0083 (13) | 0.0077 (14) |
| C5 | 0.0243 (15) | 0.0412 (16) | 0.0483 (17) | −0.0013 (12) | −0.0008 (13) | 0.0108 (14) |
| C6 | 0.0275 (16) | 0.063 (2) | 0.078 (3) | −0.0055 (16) | 0.0029 (15) | 0.0105 (18) |
| C7 | 0.0477 (19) | 0.0404 (16) | 0.0419 (17) | −0.0001 (14) | 0.0055 (14) | 0.0021 (14) |
| C8 | 0.074 (2) | 0.0470 (18) | 0.0439 (18) | −0.0052 (18) | 0.0120 (17) | 0.0008 (15) |
| C9 | 0.085 (3) | 0.097 (3) | 0.074 (3) | 0.008 (3) | 0.017 (2) | −0.022 (2) |
| C10 | 0.135 (5) | 0.103 (4) | 0.079 (3) | 0.013 (4) | 0.045 (3) | −0.028 (3) |
| C11 | 0.191 (6) | 0.051 (2) | 0.058 (3) | −0.025 (3) | 0.044 (3) | −0.005 (2) |
| C12 | 0.121 (4) | 0.100 (4) | 0.077 (3) | −0.053 (3) | 0.042 (3) | −0.038 (3) |
| C13 | 0.284 (10) | 0.077 (3) | 0.079 (3) | −0.040 (5) | 0.069 (5) | −0.028 (3) |
| N1 | 0.0341 (14) | 0.0423 (14) | 0.0448 (14) | −0.0016 (11) | −0.0021 (11) | −0.0011 (12) |
| N2 | 0.084 (2) | 0.0570 (19) | 0.0613 (19) | −0.0162 (18) | −0.0083 (17) | −0.0069 (16) |
| N3 | 0.0279 (13) | 0.0415 (14) | 0.0459 (13) | −0.0002 (10) | 0.0022 (11) | 0.0015 (11) |
| N4 | 0.0383 (14) | 0.0470 (15) | 0.0480 (14) | 0.0015 (12) | 0.0099 (12) | −0.0030 (12) |
| N5 | 0.183 (5) | 0.101 (3) | 0.079 (3) | −0.074 (3) | 0.053 (3) | −0.042 (2) |
| O1 | 0.0351 (12) | 0.0517 (12) | 0.0495 (12) | −0.0071 (10) | 0.0019 (9) | −0.0078 (10) |
Geometric parameters (Å, º) top
| Ni1—N1 | 2.095 (2) | C6—H6B | 0.9600 |
| Ni1—N1i | 2.095 (2) | C6—H6C | 0.9600 |
| Ni1—N3 | 1.984 (2) | C7—C8 | 1.480 (4) |
| Ni1—N3i | 1.984 (2) | C7—N4 | 1.333 (4) |
| Ni1—O1i | 2.069 (2) | C7—O1 | 1.275 (3) |
| Ni1—O1 | 2.069 (2) | C8—C9 | 1.376 (5) |
| C1—H1 | 0.9300 | C8—C12 | 1.367 (6) |
| C1—C2 | 1.374 (5) | C9—H9 | 0.9300 |
| C1—N1 | 1.316 (4) | C9—C10 | 1.381 (6) |
| C2—H2 | 0.9300 | C10—H10 | 0.9300 |
| C2—N2 | 1.322 (5) | C10—C11 | 1.350 (7) |
| C3—H3 | 0.9300 | C11—C13 | 1.515 (6) |
| C3—C4 | 1.382 (4) | C11—N5 | 1.295 (7) |
| C3—N2 | 1.328 (4) | C12—H12 | 0.9300 |
| C4—C5 | 1.474 (4) | C12—N5 | 1.336 (5) |
| C4—N1 | 1.354 (4) | C13—H13A | 0.9600 |
| C5—C6 | 1.493 (4) | C13—H13B | 0.9600 |
| C5—N3 | 1.288 (4) | C13—H13C | 0.9600 |
| C6—H6A | 0.9600 | N3—N4 | 1.370 (3) |
| | | |
| N1—Ni1—N1i | 92.07 (13) | H6B—C6—H6C | 109.5 |
| N3i—Ni1—N1 | 103.09 (10) | N4—C7—C8 | 115.9 (3) |
| N3—Ni1—N1 | 78.43 (10) | O1—C7—C8 | 118.1 (3) |
| N3—Ni1—N1i | 103.09 (10) | O1—C7—N4 | 126.0 (3) |
| N3i—Ni1—N1i | 78.43 (10) | C9—C8—C7 | 123.4 (3) |
| N3i—Ni1—N3 | 177.86 (13) | C12—C8—C7 | 120.8 (3) |
| N3i—Ni1—O1i | 76.86 (9) | C12—C8—C9 | 115.8 (4) |
| N3—Ni1—O1i | 101.68 (9) | C8—C9—H9 | 120.6 |
| N3i—Ni1—O1 | 101.68 (9) | C8—C9—C10 | 118.8 (5) |
| N3—Ni1—O1 | 76.86 (9) | C10—C9—H9 | 120.6 |
| O1—Ni1—N1i | 91.00 (9) | C9—C10—H10 | 119.7 |
| O1i—Ni1—N1i | 155.16 (8) | C11—C10—C9 | 120.5 (5) |
| O1—Ni1—N1 | 155.16 (8) | C11—C10—H10 | 119.7 |
| O1i—Ni1—N1 | 91.00 (9) | C10—C11—C13 | 121.2 (5) |
| O1i—Ni1—O1 | 96.45 (13) | N5—C11—C10 | 121.8 (4) |
| C2—C1—H1 | 119.2 | N5—C11—C13 | 117.0 (6) |
| N1—C1—H1 | 119.2 | C8—C12—H12 | 117.5 |
| N1—C1—C2 | 121.7 (3) | N5—C12—C8 | 124.9 (5) |
| C1—C2—H2 | 119.0 | N5—C12—H12 | 117.5 |
| N2—C2—C1 | 122.1 (4) | C11—C13—H13A | 109.5 |
| N2—C2—H2 | 119.0 | C11—C13—H13B | 109.5 |
| C4—C3—H3 | 118.5 | C11—C13—H13C | 109.5 |
| N2—C3—H3 | 118.5 | H13A—C13—H13B | 109.5 |
| N2—C3—C4 | 122.9 (3) | H13A—C13—H13C | 109.5 |
| C3—C4—C5 | 124.7 (3) | H13B—C13—H13C | 109.5 |
| N1—C4—C3 | 119.3 (3) | C1—N1—Ni1 | 130.3 (2) |
| N1—C4—C5 | 116.0 (3) | C1—N1—C4 | 117.6 (3) |
| C4—C5—C6 | 122.4 (3) | C4—N1—Ni1 | 112.00 (19) |
| N3—C5—C4 | 113.4 (3) | C2—N2—C3 | 116.3 (3) |
| N3—C5—C6 | 124.2 (3) | C5—N3—Ni1 | 119.8 (2) |
| C5—C6—H6A | 109.5 | C5—N3—N4 | 121.2 (3) |
| C5—C6—H6B | 109.5 | N4—N3—Ni1 | 118.93 (19) |
| C5—C6—H6C | 109.5 | C7—N4—N3 | 107.9 (2) |
| H6A—C6—H6B | 109.5 | C11—N5—C12 | 118.1 (5) |
| H6A—C6—H6C | 109.5 | C7—O1—Ni1 | 110.16 (18) |
| | | |
| Ni1—N3—N4—C7 | 2.4 (3) | N1i—Ni1—N3—N4 | −88.3 (2) |
| C1—C2—N2—C3 | −0.6 (6) | N1—Ni1—N3—N4 | −177.7 (2) |
| C2—C1—N1—Ni1 | −179.0 (3) | N1i—Ni1—O1—C7 | 101.3 (2) |
| C2—C1—N1—C4 | −0.6 (5) | N1—Ni1—O1—C7 | 4.2 (3) |
| C3—C4—C5—C6 | 6.7 (5) | N1—C1—C2—N2 | 0.9 (6) |
| C3—C4—C5—N3 | −174.0 (3) | N1—C4—C5—C6 | −175.0 (3) |
| C3—C4—N1—Ni1 | 178.7 (2) | N1—C4—C5—N3 | 4.3 (4) |
| C3—C4—N1—C1 | 0.0 (4) | N2—C3—C4—C5 | 178.5 (3) |
| C4—C3—N2—C2 | 0.0 (5) | N2—C3—C4—N1 | 0.3 (5) |
| C4—C5—N3—Ni1 | −7.2 (3) | N3—Ni1—N1—C1 | 175.5 (3) |
| C4—C5—N3—N4 | 176.5 (2) | N3i—Ni1—N1—C1 | −6.1 (3) |
| C5—C4—N1—Ni1 | 0.3 (3) | N3—Ni1—N1—C4 | −2.99 (19) |
| C5—C4—N1—C1 | −178.3 (3) | N3i—Ni1—N1—C4 | 175.47 (19) |
| C5—N3—N4—C7 | 178.7 (3) | N3i—Ni1—N3—C5 | −129.6 (2) |
| C6—C5—N3—Ni1 | 172.0 (2) | N3i—Ni1—N3—N4 | 46.8 (2) |
| C6—C5—N3—N4 | −4.3 (4) | N3i—Ni1—O1—C7 | 179.7 (2) |
| C7—C8—C9—C10 | 178.5 (4) | N3—Ni1—O1—C7 | −1.9 (2) |
| C7—C8—C12—N5 | −179.1 (5) | N4—C7—C8—C9 | −6.0 (5) |
| C8—C7—N4—N3 | 176.7 (2) | N4—C7—C8—C12 | 173.5 (4) |
| C8—C7—O1—Ni1 | −176.9 (2) | N4—C7—O1—Ni1 | 4.4 (4) |
| C8—C9—C10—C11 | 1.4 (8) | O1—Ni1—N1—C1 | 169.4 (3) |
| C8—C12—N5—C11 | −0.2 (9) | O1i—Ni1—N1—C1 | −82.8 (3) |
| C9—C8—C12—N5 | 0.5 (8) | O1—Ni1—N1—C4 | −9.0 (3) |
| C9—C10—C11—C13 | 179.2 (5) | O1i—Ni1—N1—C4 | 98.7 (2) |
| C9—C10—C11—N5 | −1.1 (9) | O1i—Ni1—N3—C5 | −82.7 (2) |
| C10—C11—N5—C12 | 0.5 (9) | O1—Ni1—N3—C5 | −176.7 (2) |
| C12—C8—C9—C10 | −1.0 (7) | O1—Ni1—N3—N4 | −0.36 (19) |
| C13—C11—N5—C12 | −179.8 (5) | O1i—Ni1—N3—N4 | 93.6 (2) |
| N1i—Ni1—N1—C1 | 72.5 (3) | O1i—Ni1—O1—C7 | −102.4 (2) |
| N1i—Ni1—N1—C4 | −105.9 (2) | O1—C7—C8—C9 | 175.2 (4) |
| N1i—Ni1—N3—C5 | 95.3 (2) | O1—C7—C8—C12 | −5.3 (5) |
| N1—Ni1—N3—C5 | 5.9 (2) | O1—C7—N4—N3 | −4.6 (4) |
| Symmetry code: (i) −x, y, −z+3/2. |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| C2—H2···N5ii | 0.93 | 2.54 | 3.448 (6) | 164 |
| C3—H3···O1iii | 0.93 | 2.38 | 3.278 (4) | 161 |
| Symmetry codes: (ii) −x, y+1, −z+3/2; (iii) x+1/2, y+1/2, −z+3/2. |
Di-µ-azido-
κ4N1:
N1-bis({6-methyl-
N'-[1-(pyrazin-2-yl-
κN1)ethylidene]nicotinohydrazidato-
κ2N',
O}nickel(II)) (II)
top
Crystal data top
| [Cu2(C13H12N5O)2(N3)2] | Z = 1 |
| Mr = 719.71 | F(000) = 366 |
| Triclinic, P1 | Dx = 1.638 Mg m−3 |
| a = 7.8491 (5) Å | Mo Kα radiation, λ = 0.71073 Å |
| b = 7.8762 (6) Å | Cell parameters from 3711 reflections |
| c = 12.3391 (8) Å | θ = 2.7–26.4° |
| α = 100.236 (2)° | µ = 1.51 mm−1 |
| β = 94.225 (2)° | T = 293 K |
| γ = 101.865 (2)° | Block, blue |
| V = 729.81 (9) Å3 | 0.24 × 0.2 × 0.17 mm |
Data collection top
CCD area detector
diffractometer | 3163 independent reflections |
| Radiation source: fine-focus sealed tube | 2438 reflections with I > 2σ(I) |
| Graphite monochromator | Rint = 0.034 |
| phi and ω scans | θmax = 27.1°, θmin = 2.7° |
Absorption correction: multi-scan
(CrysAlis PRO; Agilent, 2014) | h = −10→8 |
| Tmin = 0.713, Tmax = 0.783 | k = −10→10 |
| 7758 measured reflections | l = −15→15 |
Refinement top
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.039 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.086 | H atoms treated by a mixture of independent and constrained refinement |
| S = 1.03 | w = 1/[σ2(Fo2) + (0.0317P)2 + 0.4086P]
where P = (Fo2 + 2Fc2)/3 |
| 3163 reflections | (Δ/σ)max < 0.001 |
| 210 parameters | Δρmax = 0.43 e Å−3 |
| 0 restraints | Δρmin = −0.32 e Å−3 |
Special details top
Geometry. All esds (except the esd in the dihedral angle between two l.s. planes)
are estimated using the full covariance matrix. The cell esds are taken
into account individually in the estimation of esds in distances, angles
and torsion angles; correlations between esds in cell parameters are only
used when they are defined by crystal symmetry. An approximate (isotropic)
treatment of cell esds is used for estimating esds involving l.s. planes. |
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and
goodness of fit S are based on F2, conventional R-factors R are based
on F, with F set to zero for negative F2. The threshold expression of
F2 > 2sigma(F2) is used only for calculating R-factors(gt) etc. and is
not relevant to the choice of reflections for refinement. R-factors based
on F2 are statistically about twice as large as those based on F, and R-
factors based on ALL data will be even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| Cu1 | 0.48975 (4) | 0.69878 (4) | 0.00180 (3) | 0.03177 (13) | |
| C1 | 0.4083 (4) | 0.5429 (4) | −0.2461 (2) | 0.0435 (7) | |
| H1 | 0.3211 | 0.4542 | −0.2304 | 0.052* | |
| C2 | 0.4464 (5) | 0.5393 (4) | −0.3541 (2) | 0.0538 (9) | |
| H2 | 0.3855 | 0.4453 | −0.4091 | 0.065* | |
| C3 | 0.6506 (4) | 0.7945 (4) | −0.3010 (2) | 0.0444 (7) | |
| H3 | 0.7336 | 0.8856 | −0.3181 | 0.053* | |
| C4 | 0.6197 (3) | 0.8001 (3) | −0.1912 (2) | 0.0323 (6) | |
| C5 | 0.7154 (3) | 0.9364 (3) | −0.0967 (2) | 0.0302 (6) | |
| C6 | 0.8551 (4) | 1.0884 (4) | −0.1088 (2) | 0.0411 (7) | |
| H6A | 0.8903 | 1.1675 | −0.0384 | 0.062* | |
| H6B | 0.8112 | 1.1500 | −0.1613 | 0.062* | |
| H6C | 0.9540 | 1.0459 | −0.1346 | 0.062* | |
| C7 | 0.6608 (3) | 0.9474 (3) | 0.1760 (2) | 0.0306 (6) | |
| C8 | 0.7331 (3) | 1.0384 (3) | 0.2907 (2) | 0.0316 (6) | |
| C9 | 0.8744 (4) | 1.1820 (4) | 0.3129 (2) | 0.0398 (7) | |
| H9 | 0.9230 | 1.2282 | 0.2550 | 0.048* | |
| C10 | 0.9423 (4) | 1.2556 (4) | 0.4195 (2) | 0.0428 (7) | |
| H10 | 1.0378 | 1.3516 | 0.4346 | 0.051* | |
| C11 | 0.8694 (4) | 1.1875 (4) | 0.5048 (2) | 0.0452 (8) | |
| C12 | 0.6652 (4) | 0.9781 (4) | 0.3811 (2) | 0.0442 (7) | |
| H12 | 0.5691 | 0.8828 | 0.3680 | 0.053* | |
| C13 | 0.9416 (5) | 1.2654 (5) | 0.6235 (3) | 0.0721 (12) | |
| H13A | 0.8499 | 1.2987 | 0.6641 | 0.108* | |
| H13B | 1.0337 | 1.3679 | 0.6266 | 0.108* | |
| H13C | 0.9873 | 1.1792 | 0.6557 | 0.108* | |
| N1 | 0.4956 (3) | 0.6724 (3) | −0.16489 (17) | 0.0331 (5) | |
| N2 | 0.5656 (4) | 0.6634 (4) | −0.3827 (2) | 0.0540 (7) | |
| N3 | 0.6642 (3) | 0.9067 (3) | −0.00367 (17) | 0.0286 (5) | |
| N4 | 0.7384 (3) | 1.0147 (3) | 0.09616 (17) | 0.0319 (5) | |
| N5 | 0.7310 (4) | 1.0502 (4) | 0.4864 (2) | 0.0530 (7) | |
| N6 | 0.6952 (3) | 0.5074 (3) | 0.00231 (19) | 0.0349 (5) | |
| N7 | 0.7937 (3) | 0.5260 (3) | −0.0673 (2) | 0.0373 (6) | |
| N8 | 0.8893 (4) | 0.5527 (4) | −0.1304 (3) | 0.0647 (8) | |
| O1 | 0.5340 (2) | 0.8106 (2) | 0.16055 (14) | 0.0360 (4) | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| Cu1 | 0.0343 (2) | 0.0281 (2) | 0.0298 (2) | 0.00161 (13) | 0.00432 (14) | 0.00316 (13) |
| C1 | 0.0509 (18) | 0.0366 (17) | 0.0368 (17) | 0.0012 (14) | −0.0028 (14) | 0.0043 (13) |
| C2 | 0.075 (2) | 0.0443 (19) | 0.0319 (17) | 0.0039 (17) | −0.0035 (16) | −0.0028 (14) |
| C3 | 0.0541 (19) | 0.0433 (18) | 0.0345 (16) | 0.0070 (15) | 0.0100 (14) | 0.0068 (14) |
| C4 | 0.0353 (15) | 0.0323 (15) | 0.0317 (15) | 0.0120 (12) | 0.0039 (12) | 0.0077 (12) |
| C5 | 0.0315 (14) | 0.0292 (14) | 0.0321 (15) | 0.0096 (12) | 0.0056 (12) | 0.0076 (11) |
| C6 | 0.0409 (17) | 0.0422 (17) | 0.0373 (16) | 0.0014 (14) | 0.0080 (13) | 0.0078 (13) |
| C7 | 0.0287 (14) | 0.0290 (14) | 0.0334 (15) | 0.0089 (12) | 0.0037 (12) | 0.0015 (12) |
| C8 | 0.0348 (15) | 0.0304 (14) | 0.0294 (14) | 0.0078 (12) | 0.0052 (12) | 0.0041 (11) |
| C9 | 0.0448 (17) | 0.0396 (17) | 0.0330 (16) | 0.0028 (14) | 0.0075 (13) | 0.0079 (13) |
| C10 | 0.0446 (17) | 0.0394 (17) | 0.0342 (16) | −0.0079 (13) | −0.0018 (14) | 0.0032 (13) |
| C11 | 0.0533 (19) | 0.0441 (18) | 0.0343 (16) | 0.0047 (15) | −0.0003 (14) | 0.0068 (14) |
| C12 | 0.0480 (18) | 0.0410 (17) | 0.0393 (17) | 0.0000 (14) | 0.0060 (14) | 0.0081 (14) |
| C13 | 0.094 (3) | 0.071 (3) | 0.0342 (18) | −0.011 (2) | −0.0101 (19) | 0.0060 (17) |
| N1 | 0.0375 (13) | 0.0313 (12) | 0.0290 (12) | 0.0070 (10) | −0.0014 (10) | 0.0048 (10) |
| N2 | 0.0734 (19) | 0.0508 (17) | 0.0317 (14) | 0.0052 (15) | 0.0070 (13) | 0.0014 (12) |
| N3 | 0.0323 (12) | 0.0262 (11) | 0.0262 (11) | 0.0063 (9) | 0.0038 (10) | 0.0020 (9) |
| N4 | 0.0338 (12) | 0.0320 (12) | 0.0277 (12) | 0.0041 (10) | 0.0048 (10) | 0.0030 (10) |
| N5 | 0.0651 (18) | 0.0545 (17) | 0.0343 (15) | 0.0002 (14) | 0.0059 (13) | 0.0105 (13) |
| N6 | 0.0328 (13) | 0.0313 (13) | 0.0384 (13) | 0.0020 (10) | 0.0054 (11) | 0.0064 (10) |
| N7 | 0.0323 (13) | 0.0303 (13) | 0.0465 (15) | 0.0032 (10) | 0.0026 (12) | 0.0059 (11) |
| N8 | 0.0599 (19) | 0.064 (2) | 0.075 (2) | 0.0091 (15) | 0.0360 (17) | 0.0213 (16) |
| O1 | 0.0375 (11) | 0.0334 (10) | 0.0321 (10) | −0.0017 (8) | 0.0063 (9) | 0.0036 (8) |
Geometric parameters (Å, º) top
| Cu1—Cu1i | 3.1610 (7) | C7—C8 | 1.481 (4) |
| Cu1—N1 | 2.035 (2) | C7—N4 | 1.328 (3) |
| Cu1—N3 | 1.921 (2) | C7—O1 | 1.283 (3) |
| Cu1—N6i | 1.933 (2) | C8—C9 | 1.383 (4) |
| Cu1—N6 | 2.423 (2) | C8—C12 | 1.388 (4) |
| Cu1—O1 | 1.9733 (18) | C9—H9 | 0.9300 |
| C1—H1 | 0.9300 | C9—C10 | 1.359 (4) |
| C1—C2 | 1.384 (4) | C10—H10 | 0.9300 |
| C1—N1 | 1.328 (3) | C10—C11 | 1.373 (4) |
| C2—H2 | 0.9300 | C11—C13 | 1.503 (4) |
| C2—N2 | 1.323 (4) | C11—N5 | 1.340 (4) |
| C3—H3 | 0.9300 | C12—H12 | 0.9300 |
| C3—C4 | 1.388 (4) | C12—N5 | 1.340 (4) |
| C3—N2 | 1.332 (4) | C13—H13A | 0.9600 |
| C4—C5 | 1.474 (4) | C13—H13B | 0.9600 |
| C4—N1 | 1.351 (3) | C13—H13C | 0.9600 |
| C5—C6 | 1.485 (4) | N3—N4 | 1.378 (3) |
| C5—N3 | 1.288 (3) | N6—Cu1i | 1.933 (2) |
| C6—H6A | 0.9600 | N6—N7 | 1.205 (3) |
| C6—H6B | 0.9600 | N7—N8 | 1.137 (3) |
| C6—H6C | 0.9600 | | |
| | | |
| N1—Cu1—Cu1i | 92.99 (6) | O1—C7—C8 | 119.2 (2) |
| N1—Cu1—N6 | 88.24 (8) | O1—C7—N4 | 125.2 (2) |
| N3—Cu1—Cu1i | 132.40 (6) | C9—C8—C7 | 122.1 (2) |
| N3—Cu1—N1 | 79.57 (9) | C9—C8—C12 | 117.0 (3) |
| N3—Cu1—N6i | 175.77 (9) | C12—C8—C7 | 120.9 (2) |
| N3—Cu1—N6 | 94.82 (8) | C8—C9—H9 | 120.0 |
| N3—Cu1—O1 | 79.90 (8) | C10—C9—C8 | 120.0 (3) |
| N6—Cu1—Cu1i | 37.67 (5) | C10—C9—H9 | 120.0 |
| N6i—Cu1—Cu1i | 49.99 (7) | C9—C10—H10 | 120.1 |
| N6i—Cu1—N1 | 97.11 (9) | C9—C10—C11 | 119.7 (3) |
| N6i—Cu1—N6 | 87.66 (9) | C11—C10—H10 | 120.1 |
| N6i—Cu1—O1 | 103.19 (9) | C10—C11—C13 | 121.1 (3) |
| O1—Cu1—Cu1i | 104.09 (6) | N5—C11—C10 | 121.9 (3) |
| O1—Cu1—N1 | 159.01 (8) | N5—C11—C13 | 117.0 (3) |
| O1—Cu1—N6 | 97.79 (8) | C8—C12—H12 | 118.3 |
| C2—C1—H1 | 119.9 | N5—C12—C8 | 123.4 (3) |
| N1—C1—H1 | 119.9 | N5—C12—H12 | 118.3 |
| N1—C1—C2 | 120.3 (3) | C11—C13—H13A | 109.5 |
| C1—C2—H2 | 118.5 | C11—C13—H13B | 109.5 |
| N2—C2—C1 | 123.0 (3) | C11—C13—H13C | 109.5 |
| N2—C2—H2 | 118.5 | H13A—C13—H13B | 109.5 |
| C4—C3—H3 | 118.8 | H13A—C13—H13C | 109.5 |
| N2—C3—H3 | 118.8 | H13B—C13—H13C | 109.5 |
| N2—C3—C4 | 122.3 (3) | C1—N1—Cu1 | 129.3 (2) |
| C3—C4—C5 | 124.9 (3) | C1—N1—C4 | 118.1 (2) |
| N1—C4—C3 | 119.9 (2) | C4—N1—Cu1 | 112.23 (17) |
| N1—C4—C5 | 115.3 (2) | C2—N2—C3 | 116.4 (3) |
| C4—C5—C6 | 123.3 (2) | C5—N3—Cu1 | 120.73 (18) |
| N3—C5—C4 | 112.0 (2) | C5—N3—N4 | 122.2 (2) |
| N3—C5—C6 | 124.7 (2) | N4—N3—Cu1 | 117.00 (15) |
| C5—C6—H6A | 109.5 | C7—N4—N3 | 107.8 (2) |
| C5—C6—H6B | 109.5 | C11—N5—C12 | 118.0 (2) |
| C5—C6—H6C | 109.5 | Cu1i—N6—Cu1 | 92.34 (9) |
| H6A—C6—H6B | 109.5 | N7—N6—Cu1i | 125.40 (19) |
| H6A—C6—H6C | 109.5 | N7—N6—Cu1 | 111.79 (17) |
| H6B—C6—H6C | 109.5 | N8—N7—N6 | 176.4 (3) |
| N4—C7—C8 | 115.6 (2) | C7—O1—Cu1 | 109.78 (16) |
| | | |
| Cu1i—Cu1—N1—C1 | 44.7 (2) | N1—Cu1—N6—Cu1i | −97.19 (9) |
| Cu1i—Cu1—N1—C4 | −128.65 (17) | N1—Cu1—N6—N7 | 32.66 (19) |
| Cu1i—Cu1—N3—C5 | 80.8 (2) | N1—Cu1—O1—C7 | −17.7 (3) |
| Cu1i—Cu1—N3—N4 | −95.32 (17) | N1—C1—C2—N2 | −1.7 (5) |
| Cu1i—Cu1—N6—N7 | 129.8 (2) | N1—C4—C5—C6 | −179.3 (2) |
| Cu1i—Cu1—O1—C7 | 125.85 (16) | N1—C4—C5—N3 | 1.2 (3) |
| Cu1—N3—N4—C7 | −2.6 (3) | N2—C3—C4—C5 | 176.8 (3) |
| Cu1i—N6—N7—N8 | 167 (5) | N2—C3—C4—N1 | −2.2 (4) |
| Cu1—N6—N7—N8 | 57 (5) | N3—Cu1—N1—C1 | 177.4 (3) |
| C1—C2—N2—C3 | 0.6 (5) | N3—Cu1—N1—C4 | 3.95 (17) |
| C2—C1—N1—Cu1 | −172.2 (2) | N3—Cu1—N6—Cu1i | −176.56 (9) |
| C2—C1—N1—C4 | 0.9 (4) | N3—Cu1—N6—N7 | −46.72 (19) |
| C3—C4—C5—C6 | 1.6 (4) | N3—Cu1—O1—C7 | −5.56 (17) |
| C3—C4—C5—N3 | −177.9 (3) | N4—C7—C8—C9 | −1.2 (4) |
| C3—C4—N1—Cu1 | 175.2 (2) | N4—C7—C8—C12 | −178.7 (3) |
| C3—C4—N1—C1 | 1.0 (4) | N4—C7—O1—Cu1 | 6.6 (3) |
| C4—C3—N2—C2 | 1.4 (5) | N6—Cu1—N1—C1 | 82.1 (2) |
| C4—C5—N3—Cu1 | 2.5 (3) | N6i—Cu1—N1—C1 | −5.3 (3) |
| C4—C5—N3—N4 | 178.4 (2) | N6i—Cu1—N1—C4 | −178.70 (18) |
| C5—C4—N1—Cu1 | −3.9 (3) | N6—Cu1—N1—C4 | −91.27 (18) |
| C5—C4—N1—C1 | −178.1 (2) | N6i—Cu1—N3—C5 | −42.1 (13) |
| C5—N3—N4—C7 | −178.6 (2) | N6—Cu1—N3—C5 | 83.6 (2) |
| C6—C5—N3—Cu1 | −177.0 (2) | N6i—Cu1—N3—N4 | 141.8 (12) |
| C6—C5—N3—N4 | −1.1 (4) | N6—Cu1—N3—N4 | −92.47 (17) |
| C7—C8—C9—C10 | −176.5 (3) | N6i—Cu1—N6—Cu1i | 0.0 |
| C7—C8—C12—N5 | 176.7 (3) | N6i—Cu1—N6—N7 | 129.8 (2) |
| C8—C7—N4—N3 | 175.4 (2) | N6i—Cu1—O1—C7 | 177.39 (17) |
| C8—C7—O1—Cu1 | −171.71 (18) | N6—Cu1—O1—C7 | 87.98 (17) |
| C8—C9—C10—C11 | −0.4 (5) | O1—Cu1—N1—C1 | −170.5 (2) |
| C8—C12—N5—C11 | 0.0 (5) | O1—Cu1—N1—C4 | 16.1 (3) |
| C9—C8—C12—N5 | −0.9 (5) | O1—Cu1—N3—C5 | −179.3 (2) |
| C9—C10—C11—C13 | 179.8 (3) | O1—Cu1—N3—N4 | 4.60 (16) |
| C9—C10—C11—N5 | −0.5 (5) | O1—Cu1—N6—Cu1i | 103.00 (9) |
| C10—C11—N5—C12 | 0.7 (5) | O1—Cu1—N6—N7 | −127.16 (18) |
| C12—C8—C9—C10 | 1.1 (4) | O1—C7—C8—C9 | 177.2 (2) |
| C13—C11—N5—C12 | −179.6 (3) | O1—C7—C8—C12 | −0.3 (4) |
| N1—Cu1—N3—C5 | −3.7 (2) | O1—C7—N4—N3 | −2.9 (3) |
| N1—Cu1—N3—N4 | −179.79 (18) | | |
| Symmetry code: (i) −x+1, −y+1, −z. |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| C6—H6C···N4ii | 0.96 | 2.58 | 3.450 (4) | 150 |
| Symmetry code: (ii) −x+2, −y+2, −z. |
Inhibition of jack bean urease by HL, (I) and (II) top| Compound | IC50 (µM) | Compound | IC50 (µM) |
| (I) | 9.68±1.4 | Ni(N3)2 | 5.94±1.1 |
| (II) | 1.32±0.4 | Cu(N3)2 | 1.76±0.7 |
| HL | 105±4.8 | AHA | 7.82±1.6 |
The BSA binding constants and parameters top| Complex | KSV (M-1) | kq (M-1s-1) | K (M-1) | n |
| (I) | 3.14±0.01 × 104 | 3.14±0.01 × 1012 | 2.36±0.09 × 104 | 1.37 |
| (II) | 7.71±0.06 × 104 | 7.71±0.06 × 1012 | 1.74±0.11 × 104 | 1.23 |

 Subscribe to Acta Crystallographica Section C: Structural Chemistry
Subscribe to Acta Crystallographica Section C: Structural Chemistry
The full text of this article is available to subscribers to the journal.
If you have already registered and are using a computer listed in your registration details, please email
support@iucr.org for assistance.

 Subscribe to Acta Crystallographica Section C: Structural Chemistry
Subscribe to Acta Crystallographica Section C: Structural Chemistry
 journal menu
journal menu











