In the crystalline state, the centrosymmetric molecule 1,2,4,5-tetrakis(cyanomethyl)benzene, C
14H
10N
4, has one cyanomethyl group in the benzene plane and one cyanomethyl group rotated 67.2 (2)° out of the benzene plane. Molecules of methyl 3,4,5-triacetoxybenzoate, C
14H
14O
8, form chains with each molecule twisted
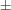
89.6 (1)° from the preceding molecule. In this orientation, a close C—H

O contact is formed, with an H

O distance of 2.34 Å. The structure of 2-(
N-phthalimidomethyl)benzoic acid, C
16H
11NO
4, reveals hydrogen-bonded dimers linked by the carboxyl groups of adjacent molecules. The O4

O3 distance is 2.636 (2) Å and the O4—H

O3 angle is 171 (2)°.
Supporting information
CCDC references: 142778; 142779; 142780
Crystals of (I) were prepared by the method of Buckland et al. (1983) and recrystallized from acetonitrile (m.p. 484–487 K, literature m.p. 486–488 K). Compound (II) can be prepared by acylation of the alcohol functions of gallic acid, followed by esterification of the carboxyl group (m.p. 404.0–405.5 K). Crystals of (III) were prepared with the use of the method of Sasaki et al. (1978) to protect 2-methylaminobenzoic acid with phthalimide (m.p. 535.0–537.0 K). Melting points were measured on a Mel-Temp melting point apparatus from Laboratory Instruments, Holliston, Massachusetts, USA.
H atoms were placed at calculated positions and refined with a riding model (Cmethyl—H = 0.98 Å; Cmethylene—H = 0.99 Å; Caromatic—H = 0.95 Å). The Uiso value for a given H atom was assigned as 1.2 times the Uiso of the atom to which it is attached (1.5 for methyl). The methyl groups in (II) were constrained to ideal tetrahedral geometries. The acidic proton H4B in (III) was refined isotropically. Data were collected at 153 K in groups of 606, 435, and 230 frames at ϕ settings of 0°, 90°, and 180°, respectively. Crystals were attached to glass fibres with a minimum of silicone cement.
Data collection: SMART (Bruker, 1997) for (I), (II); SMART version 5.054 (Bruker, 1999) for (III). Cell refinement: SMART (Bruker, 1997) for (I), (II); SMART version 5.054 (Bruker, 1999) for (III). Data reduction: SAINT-Plus (Bruker, 1997) for (I), (II); SAINT-Plus version 6.0 (Bruker, 1999) for (III). Program(s) used to solve structure: SHELXS97 (Sheldrick, 1990) for (I), (II); SHELXS99 (Sheldrick, 1990) for (III). Program(s) used to refine structure: SHELXTL/PC (Sheldrick, 1997) for (I), (II); SHELXL99 (Sheldrick, 1999) for (III). Molecular graphics: SHELXTL/PC (Sheldrick, 1997) for (I), (II); SHELXTL99 (Sheldrick, 1999) for (III). Software used to prepare material for publication: SHELXTL/PC (Sheldrick, 1997) for (I), (II); SHELXTL99 (Sheldrick, 1999) for (III).
(I) 1,2,4,5-tetrakis(cyanomethyl)benzene
top
Crystal data top
| C14H10N4 | Dx = 1.359 Mg m−3 |
| Mr = 234.26 | Melting point: 485 K |
| Monoclinic, P21/c | Mo Kα radiation, λ = 0.71073 Å |
| a = 8.818 (1) Å | Cell parameters from 853 reflections |
| b = 8.377 (1) Å | θ = 2.4–28.2° |
| c = 8.174 (1) Å | µ = 0.09 mm−1 |
| β = 108.51 (1)° | T = 153 K |
| V = 572.48 (15) Å3 | Square plate, colourless |
| Z = 2 | 0.12 × 0.11 × 0.06 mm |
| F(000) = 244 | |
Data collection top
Bruker SMART 1000 CCD
diffractometer | 1335 independent reflections |
| Radiation source: standard-focus sealed tube | 836 reflections with I > 2σ(I) |
| Graphite monochromator | Rint = 0.035 |
| ω scans | θmax = 28.2°, θmin = 2.4° |
Absorption correction: numerical
face-indexed (SHELXTL/PC; Sheldrick, 1997) | h = −6→11 |
| Tmin = 0.985, Tmax = 0.995 | k = −10→10 |
| 3540 measured reflections | l = −10→6 |
Refinement top
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.044 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.098 | H-atom parameters constrained |
| S = 1.00 | w = 1/[σ2(Fo2) + (0.04Fo2)2] |
| 1335 reflections | (Δ/σ)max < 0.001 |
| 82 parameters | Δρmax = 0.26 e Å−3 |
| 0 restraints | Δρmin = −0.16 e Å−3 |
Crystal data top
| C14H10N4 | V = 572.48 (15) Å3 |
| Mr = 234.26 | Z = 2 |
| Monoclinic, P21/c | Mo Kα radiation |
| a = 8.818 (1) Å | µ = 0.09 mm−1 |
| b = 8.377 (1) Å | T = 153 K |
| c = 8.174 (1) Å | 0.12 × 0.11 × 0.06 mm |
| β = 108.51 (1)° | |
Data collection top
Bruker SMART 1000 CCD
diffractometer | 1335 independent reflections |
Absorption correction: numerical
face-indexed (SHELXTL/PC; Sheldrick, 1997) | 836 reflections with I > 2σ(I) |
| Tmin = 0.985, Tmax = 0.995 | Rint = 0.035 |
| 3540 measured reflections | |
Refinement top
| R[F2 > 2σ(F2)] = 0.044 | 0 restraints |
| wR(F2) = 0.098 | H-atom parameters constrained |
| S = 1.00 | Δρmax = 0.26 e Å−3 |
| 1335 reflections | Δρmin = −0.16 e Å−3 |
| 82 parameters | |
Special details top
Experimental. For all compounds, the crystal to detector distance was 5.023 cm. Each frame covered −0.3° in ω for 20 s for (I), for 15 s for (II) and for 10 s for (III). Anisotropic displacement parameters were used for all non-H atoms. |
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| C1 | 0.56432 (18) | 0.57605 (17) | 0.15967 (18) | 0.0195 (4) | |
| C2 | 0.66345 (19) | 0.48312 (17) | 0.09439 (19) | 0.0197 (4) | |
| C3 | 0.59668 (19) | 0.40878 (17) | −0.06516 (18) | 0.0208 (4) | |
| H3A | 0.6634 | 0.3458 | −0.1106 | 0.025* | |
| C4 | 0.62724 (19) | 0.66513 (19) | 0.32959 (18) | 0.0243 (4) | |
| H4A | 0.7178 | 0.7335 | 0.3263 | 0.029* | |
| H4B | 0.5419 | 0.7359 | 0.3426 | 0.029* | |
| C5 | 0.6807 (2) | 0.5601 (2) | 0.4801 (2) | 0.0270 (4) | |
| C6 | 0.84030 (19) | 0.46508 (19) | 0.19484 (19) | 0.0250 (4) | |
| H6A | 0.8896 | 0.5724 | 0.2161 | 0.030* | |
| H6B | 0.8510 | 0.4164 | 0.3083 | 0.030* | |
| C7 | 0.92794 (19) | 0.36768 (19) | 0.1071 (2) | 0.0238 (4) | |
| N1 | 0.72273 (19) | 0.47705 (19) | 0.59774 (19) | 0.0400 (4) | |
| N2 | 0.99652 (16) | 0.29143 (17) | 0.03792 (17) | 0.0316 (4) | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| C1 | 0.0212 (9) | 0.0198 (8) | 0.0178 (8) | −0.0002 (7) | 0.0066 (7) | 0.0019 (6) |
| C2 | 0.0180 (9) | 0.0222 (8) | 0.0194 (7) | −0.0002 (6) | 0.0065 (7) | 0.0034 (6) |
| C3 | 0.0240 (9) | 0.0205 (8) | 0.0206 (8) | 0.0016 (7) | 0.0108 (7) | 0.0014 (6) |
| C4 | 0.0229 (9) | 0.0271 (9) | 0.0222 (8) | 0.0003 (7) | 0.0059 (7) | −0.0029 (7) |
| C5 | 0.0242 (10) | 0.0367 (10) | 0.0208 (8) | −0.0023 (8) | 0.0081 (7) | −0.0077 (8) |
| C6 | 0.0201 (9) | 0.0308 (9) | 0.0230 (8) | 0.0011 (7) | 0.0053 (7) | −0.0003 (7) |
| C7 | 0.0179 (8) | 0.0259 (8) | 0.0242 (8) | −0.0003 (7) | 0.0019 (7) | 0.0023 (7) |
| N1 | 0.0423 (11) | 0.0531 (10) | 0.0251 (8) | 0.0058 (8) | 0.0117 (7) | 0.0028 (7) |
| N2 | 0.0237 (8) | 0.0351 (8) | 0.0336 (8) | 0.0033 (7) | 0.0058 (7) | −0.0015 (7) |
Geometric parameters (Å, º) top
| C1—C3i | 1.390 (2) | C2—C6 | 1.522 (2) |
| C1—C2 | 1.397 (2) | C4—C5 | 1.464 (2) |
| C2—C3 | 1.396 (2) | C6—C7 | 1.459 (2) |
| C3—C1i | 1.390 (2) | C5—N1 | 1.149 (2) |
| C1—C4 | 1.518 (2) | C7—N2 | 1.1447 (19) |
| | | |
| C3i—C1—C2 | 119.42 (13) | C1—C2—C6 | 120.51 (13) |
| C3—C2—C1 | 118.44 (14) | C5—C4—C1 | 113.60 (12) |
| C1i—C3—C2 | 122.14 (14) | N1—C5—C4 | 179.67 (17) |
| C3i—C1—C4 | 118.40 (13) | C7—C6—C2 | 113.72 (13) |
| C2—C1—C4 | 122.16 (14) | N2—C7—C6 | 179.8 (2) |
| C3—C2—C6 | 121.05 (14) | | |
| | | |
| C2—C1—C4—C5 | −67.20 (19) | C1—C4—C5—N1 | 16 (34) |
| Symmetry code: (i) −x+1, −y+1, −z. |
(II) methyl 3,4,5-triacetoxybenzenoate
top
Crystal data top
| C14H14O8 | Dx = 1.365 Mg m−3 |
| Mr = 310.25 | Melting point = 404.0–405.5 K |
| Monoclinic, P21/n | Mo Kα radiation, λ = 0.71073 Å |
| a = 9.2233 (7) Å | Cell parameters from 4507 reflections |
| b = 11.747 (1) Å | θ = 2.3–25.5° |
| c = 13.936 (1) Å | µ = 0.11 mm−1 |
| β = 90.451 (1)° | T = 153 K |
| V = 1509.9 (2) Å3 | Truncated block, colourless |
| Z = 4 | 0.41 × 0.30 × 0.27 mm |
| F(000) = 648 | |
Data collection top
Bruker SMART 1000 CCD
diffractometer | 2789 independent reflections |
| Radiation source: standard-focus sealed tube | 2320 reflections with I > 2σ(I) |
| Graphite monochromator | Rint = 0.016 |
| ω scans | θmax = 25.5°, θmin = 2.3° |
Absorption correction: numerical
face-indexed (SHELXTL/PC; Sheldrick, 1997) | h = −11→11 |
| Tmin = 0.959, Tmax = 0.976 | k = −14→10 |
| 8088 measured reflections | l = −16→16 |
Refinement top
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.043 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.116 | H-atom parameters constrained |
| S = 1.84 | w = 1/[σ2(Fo2) + (0.04Fo2)2] |
| 2789 reflections | (Δ/σ)max = 0.008 |
| 203 parameters | Δρmax = 0.27 e Å−3 |
| 0 restraints | Δρmin = −0.26 e Å−3 |
Crystal data top
| C14H14O8 | V = 1509.9 (2) Å3 |
| Mr = 310.25 | Z = 4 |
| Monoclinic, P21/n | Mo Kα radiation |
| a = 9.2233 (7) Å | µ = 0.11 mm−1 |
| b = 11.747 (1) Å | T = 153 K |
| c = 13.936 (1) Å | 0.41 × 0.30 × 0.27 mm |
| β = 90.451 (1)° | |
Data collection top
Bruker SMART 1000 CCD
diffractometer | 2789 independent reflections |
Absorption correction: numerical
face-indexed (SHELXTL/PC; Sheldrick, 1997) | 2320 reflections with I > 2σ(I) |
| Tmin = 0.959, Tmax = 0.976 | Rint = 0.016 |
| 8088 measured reflections | |
Refinement top
| R[F2 > 2σ(F2)] = 0.043 | 0 restraints |
| wR(F2) = 0.116 | H-atom parameters constrained |
| S = 1.84 | Δρmax = 0.27 e Å−3 |
| 2789 reflections | Δρmin = −0.26 e Å−3 |
| 203 parameters | |
Special details top
Experimental. For all compounds, the crystal to detector distance was 5.023 cm. Each frame covered −0.3° in ω for 20 s for (I), for 15 s for (II) and for 10 s for (III). Anisotropic displacement parameters were used for all non-H atoms. |
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| C1 | 0.24362 (16) | 0.01512 (12) | 0.49717 (10) | 0.0264 (3) | |
| C2 | 0.25287 (16) | −0.02827 (12) | 0.40426 (10) | 0.0286 (3) | |
| H2A | 0.3222 | −0.0853 | 0.3893 | 0.034* | |
| C3 | 0.16064 (16) | 0.01251 (12) | 0.33484 (10) | 0.0284 (3) | |
| C4 | 0.05958 (15) | 0.09620 (12) | 0.35551 (10) | 0.0265 (3) | |
| C5 | 0.05181 (15) | 0.13963 (12) | 0.44761 (10) | 0.0256 (3) | |
| C6 | 0.14299 (15) | 0.09897 (12) | 0.51927 (10) | 0.0262 (3) | |
| H6A | 0.1368 | 0.1280 | 0.5827 | 0.031* | |
| C7 | 0.34029 (17) | −0.03716 (13) | 0.57130 (11) | 0.0320 (4) | |
| C8 | 0.4074 (2) | −0.04109 (17) | 0.73536 (13) | 0.0515 (5) | |
| H8A | 0.3743 | −0.0108 | 0.7969 | 0.077* | |
| H8B | 0.5103 | −0.0229 | 0.7270 | 0.077* | |
| H8C | 0.3945 | −0.1239 | 0.7344 | 0.077* | |
| C9 | 0.09629 (17) | −0.12726 (13) | 0.22126 (11) | 0.0346 (4) | |
| C10 | 0.1143 (2) | −0.16000 (16) | 0.11909 (12) | 0.0446 (4) | |
| H10A | 0.0646 | −0.2324 | 0.1070 | 0.067* | |
| H10B | 0.2177 | −0.1684 | 0.1051 | 0.067* | |
| H10C | 0.0726 | −0.1008 | 0.0778 | 0.067* | |
| C11 | 0.00377 (18) | 0.20723 (13) | 0.22017 (11) | 0.0340 (4) | |
| C12 | −0.1041 (2) | 0.21837 (16) | 0.14101 (13) | 0.0492 (5) | |
| H12A | −0.0655 | 0.2687 | 0.0913 | 0.074* | |
| H12B | −0.1942 | 0.2506 | 0.1661 | 0.074* | |
| H12C | −0.1237 | 0.1432 | 0.1134 | 0.074* | |
| C13 | −0.03401 (18) | 0.31419 (13) | 0.51374 (11) | 0.0330 (4) | |
| C14 | −0.1729 (2) | 0.37179 (15) | 0.53903 (14) | 0.0492 (5) | |
| H14A | −0.1970 | 0.3548 | 0.6059 | 0.074* | |
| H14B | −0.2506 | 0.3442 | 0.4967 | 0.074* | |
| H14C | −0.1621 | 0.4542 | 0.5311 | 0.074* | |
| O1 | 0.32352 (13) | 0.00964 (10) | 0.65800 (7) | 0.0393 (3) | |
| O2 | 0.42358 (13) | −0.11383 (10) | 0.55515 (8) | 0.0441 (3) | |
| O3 | 0.16832 (12) | −0.02718 (9) | 0.24024 (7) | 0.0339 (3) | |
| O4 | 0.03011 (14) | −0.17685 (11) | 0.28123 (8) | 0.0510 (4) | |
| O5 | −0.04070 (11) | 0.12782 (9) | 0.28506 (7) | 0.0329 (3) | |
| O6 | 0.11505 (15) | 0.25684 (12) | 0.22943 (10) | 0.0599 (4) | |
| O7 | −0.06210 (11) | 0.21413 (9) | 0.46635 (7) | 0.0313 (3) | |
| O8 | 0.08541 (13) | 0.34788 (9) | 0.53050 (8) | 0.0402 (3) | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| C1 | 0.0262 (8) | 0.0261 (7) | 0.0269 (8) | −0.0037 (6) | 0.0002 (6) | 0.0023 (6) |
| C2 | 0.0286 (8) | 0.0254 (8) | 0.0319 (8) | −0.0003 (6) | 0.0051 (6) | 0.0006 (6) |
| C3 | 0.0326 (8) | 0.0296 (8) | 0.0230 (7) | −0.0075 (6) | 0.0036 (6) | −0.0028 (6) |
| C4 | 0.0259 (8) | 0.0279 (8) | 0.0258 (7) | −0.0054 (6) | −0.0020 (6) | 0.0038 (6) |
| C5 | 0.0235 (8) | 0.0220 (7) | 0.0312 (8) | −0.0026 (5) | 0.0012 (6) | 0.0008 (6) |
| C6 | 0.0271 (8) | 0.0268 (8) | 0.0248 (7) | −0.0045 (6) | 0.0006 (6) | 0.0001 (6) |
| C7 | 0.0315 (9) | 0.0323 (9) | 0.0321 (9) | −0.0016 (7) | 0.0013 (7) | 0.0050 (6) |
| C8 | 0.0514 (11) | 0.0702 (13) | 0.0329 (9) | 0.0059 (10) | −0.0069 (8) | 0.0175 (9) |
| C9 | 0.0350 (9) | 0.0360 (9) | 0.0328 (9) | −0.0057 (7) | 0.0042 (7) | −0.0058 (7) |
| C10 | 0.0499 (11) | 0.0519 (11) | 0.0321 (9) | −0.0128 (8) | 0.0058 (8) | −0.0122 (8) |
| C11 | 0.0408 (10) | 0.0297 (8) | 0.0313 (8) | −0.0004 (7) | −0.0031 (7) | 0.0041 (6) |
| C12 | 0.0567 (12) | 0.0489 (11) | 0.0418 (10) | −0.0029 (9) | −0.0148 (9) | 0.0116 (8) |
| C13 | 0.0386 (9) | 0.0261 (8) | 0.0343 (8) | 0.0014 (7) | 0.0044 (7) | 0.0028 (6) |
| C14 | 0.0438 (11) | 0.0377 (10) | 0.0664 (12) | 0.0089 (8) | 0.0121 (9) | −0.0043 (8) |
| O1 | 0.0445 (7) | 0.0473 (7) | 0.0260 (6) | 0.0055 (5) | −0.0040 (5) | 0.0059 (5) |
| O2 | 0.0415 (7) | 0.0455 (7) | 0.0453 (7) | 0.0139 (6) | −0.0026 (5) | 0.0056 (5) |
| O3 | 0.0435 (7) | 0.0339 (6) | 0.0243 (5) | −0.0088 (5) | 0.0045 (5) | −0.0042 (4) |
| O4 | 0.0666 (9) | 0.0484 (7) | 0.0382 (7) | −0.0254 (6) | 0.0172 (6) | −0.0125 (6) |
| O5 | 0.0325 (6) | 0.0373 (6) | 0.0288 (6) | −0.0068 (5) | −0.0068 (5) | 0.0077 (4) |
| O6 | 0.0539 (9) | 0.0628 (9) | 0.0628 (9) | −0.0273 (7) | −0.0194 (7) | 0.0331 (7) |
| O7 | 0.0265 (6) | 0.0282 (6) | 0.0392 (6) | 0.0015 (4) | −0.0004 (5) | −0.0023 (4) |
| O8 | 0.0385 (7) | 0.0319 (6) | 0.0502 (7) | −0.0032 (5) | −0.0013 (5) | −0.0056 (5) |
Geometric parameters (Å, º) top
| C1—C2 | 1.395 (2) | C3—O3 | 1.4006 (17) |
| C2—C3 | 1.370 (2) | C4—O5 | 1.3939 (17) |
| C3—C4 | 1.387 (2) | C5—O7 | 1.3937 (17) |
| C4—C5 | 1.383 (2) | C7—O1 | 1.3375 (19) |
| C5—C6 | 1.386 (2) | C8—O1 | 1.4504 (19) |
| C1—C6 | 1.390 (2) | C9—O3 | 1.3751 (18) |
| C1—C7 | 1.492 (2) | C9—C10 | 1.485 (2) |
| C7—O2 | 1.2061 (18) | C11—O5 | 1.3646 (18) |
| C9—O4 | 1.1910 (19) | C11—C12 | 1.485 (2) |
| C11—O6 | 1.186 (2) | C13—O7 | 1.3719 (19) |
| C13—O8 | 1.1918 (19) | C13—C14 | 1.493 (2) |
| | | |
| C6—C1—C2 | 120.71 (14) | O2—C7—C1 | 123.81 (14) |
| C6—C1—C7 | 122.35 (13) | O1—C7—C1 | 112.56 (13) |
| C2—C1—C7 | 116.86 (13) | O4—C9—O3 | 122.19 (14) |
| C3—C2—C1 | 119.05 (14) | O4—C9—C10 | 127.43 (15) |
| C2—C3—C4 | 121.06 (13) | O3—C9—C10 | 110.37 (13) |
| C2—C3—O3 | 120.84 (14) | O6—C11—O5 | 121.87 (14) |
| C4—C3—O3 | 118.08 (13) | O6—C11—C12 | 127.67 (15) |
| C5—C4—C3 | 119.61 (13) | O5—C11—C12 | 110.46 (14) |
| C5—C4—O5 | 121.08 (13) | O8—C13—O7 | 123.34 (14) |
| C3—C4—O5 | 119.08 (13) | O8—C13—C14 | 126.61 (15) |
| C4—C5—C6 | 120.40 (13) | O7—C13—C14 | 110.05 (14) |
| C4—C5—O7 | 116.75 (13) | C7—O1—C8 | 115.96 (13) |
| C6—C5—O7 | 122.38 (13) | C9—O3—C3 | 115.95 (11) |
| C5—C6—C1 | 119.16 (13) | C11—O5—C4 | 116.60 (11) |
| O2—C7—O1 | 123.63 (14) | C13—O7—C5 | 119.24 (12) |
| | | |
| C2—C3—O3—C9 | 83.22 (17) | C4—C5—O7—C13 | −133.03 (14) |
| C3—C4—O5—C11 | −84.37 (17) | C6—C1—C7—O1 | −3.2 (2) |
Crystal data top
| C16H11NO4 | Dx = 1.464 Mg m−3 |
| Mr = 281.26 | Melting point = 262.0–264.0 K |
| Monoclinic, P21/c | Mo Kα radiation, λ = 0.71073 Å |
| a = 12.125 (1) Å | Cell parameters from 2323 reflections |
| b = 13.720 (1) Å | θ = 1.7–25.5° |
| c = 7.7247 (8) Å | µ = 0.11 mm−1 |
| β = 96.722 (2)° | T = 153 K |
| V = 1276.2 (2) Å3 | Prism, colourless |
| Z = 4 | 0.33 × 0.17 × 0.08 mm |
| F(000) = 584 | |
Data collection top
Bruker Smart 1000 CCD
diffractometer | 2365 independent reflections |
| Radiation source: standard-focus sealed tube | 1722 reflections with I > 2σ(I) |
| Graphite monochromator | Rint = 0.026 |
| ω scans | θmax = 25.5°, θmin = 1.7° |
Absorption correction: numerical
face indexed (Sheldrick, 1999) | h = −14→11 |
| Tmin = 0.974, Tmax = 0.992 | k = −16→15 |
| 6929 measured reflections | l = −9→9 |
Refinement top
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.036 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.089 | H atoms; see below |
| S = 1.10 |
w = 1/[σ2(Fo2) + (0.04Fo2)2] |
| 2365 reflections | (Δ/σ)max = 0.002 |
| 194 parameters | Δρmax = 0.16 e Å−3 |
| 0 restraints | Δρmin = −0.17 e Å−3 |
Crystal data top
| C16H11NO4 | V = 1276.2 (2) Å3 |
| Mr = 281.26 | Z = 4 |
| Monoclinic, P21/c | Mo Kα radiation |
| a = 12.125 (1) Å | µ = 0.11 mm−1 |
| b = 13.720 (1) Å | T = 153 K |
| c = 7.7247 (8) Å | 0.33 × 0.17 × 0.08 mm |
| β = 96.722 (2)° | |
Data collection top
Bruker Smart 1000 CCD
diffractometer | 2365 independent reflections |
Absorption correction: numerical
face indexed (Sheldrick, 1999) | 1722 reflections with I > 2σ(I) |
| Tmin = 0.974, Tmax = 0.992 | Rint = 0.026 |
| 6929 measured reflections | |
Refinement top
| R[F2 > 2σ(F2)] = 0.036 | 0 restraints |
| wR(F2) = 0.089 | H atoms; see below |
| S = 1.10 | Δρmax = 0.16 e Å−3 |
| 2365 reflections | Δρmin = −0.17 e Å−3 |
| 194 parameters | |
Special details top
Experimental. For all compounds, the crystal to detector distance was 5.023 cm. Each frame covered −0.3° in ω for 20 s for (I), for 15 s for (II) and for 10 s for (III). Anisotropic displacement parameters were used for all non-H atoms. |
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| N1 | 0.33447 (10) | 0.89546 (9) | 0.04519 (16) | 0.0298 (3) | |
| O1 | 0.36585 (9) | 0.72978 (8) | 0.03783 (15) | 0.0426 (3) | |
| O2 | 0.35012 (8) | 1.06013 (8) | 0.10732 (15) | 0.0379 (3) | |
| O3 | 0.09375 (9) | 0.95383 (8) | −0.35903 (14) | 0.0379 (3) | |
| O4 | −0.08878 (9) | 0.93331 (9) | −0.36400 (16) | 0.0411 (3) | |
| H4B | −0.0886 (17) | 0.9689 (17) | −0.470 (3) | 0.089 (8)* | |
| C1 | 0.48849 (11) | 0.94178 (12) | 0.22925 (19) | 0.0303 (4) | |
| C2 | 0.56875 (12) | 0.99243 (13) | 0.3340 (2) | 0.0363 (4) | |
| H2A | 0.5649 | 1.0612 | 0.3466 | 0.044* | |
| C3 | 0.65545 (13) | 0.93882 (15) | 0.4204 (2) | 0.0435 (5) | |
| H3A | 0.7119 | 0.9712 | 0.4946 | 0.052* | |
| C4 | 0.66073 (13) | 0.83854 (15) | 0.3998 (2) | 0.0459 (5) | |
| H4A | 0.7210 | 0.8036 | 0.4603 | 0.055* | |
| C5 | 0.58008 (13) | 0.78766 (13) | 0.2928 (2) | 0.0404 (4) | |
| H5A | 0.5842 | 0.7190 | 0.2783 | 0.049* | |
| C6 | 0.49380 (12) | 0.84173 (12) | 0.2085 (2) | 0.0318 (4) | |
| C7 | 0.39433 (13) | 0.81023 (12) | 0.0895 (2) | 0.0324 (4) | |
| C8 | 0.38611 (12) | 0.97760 (11) | 0.1248 (2) | 0.0294 (4) | |
| C9 | 0.23565 (12) | 0.90143 (11) | −0.0805 (2) | 0.0308 (4) | |
| H9A | 0.2431 | 0.8546 | −0.1759 | 0.037* | |
| H9B | 0.2313 | 0.9676 | −0.1319 | 0.037* | |
| C10 | 0.12754 (12) | 0.88038 (10) | −0.0048 (2) | 0.0280 (4) | |
| C11 | 0.12953 (13) | 0.84994 (11) | 0.1677 (2) | 0.0335 (4) | |
| H11A | 0.1989 | 0.8423 | 0.2373 | 0.040* | |
| C12 | 0.03246 (13) | 0.83054 (11) | 0.2396 (2) | 0.0364 (4) | |
| H12A | 0.0357 | 0.8109 | 0.3581 | 0.044* | |
| C13 | −0.06892 (13) | 0.83963 (11) | 0.1396 (2) | 0.0382 (4) | |
| H13A | −0.1354 | 0.8250 | 0.1883 | 0.046* | |
| C14 | −0.07375 (13) | 0.87000 (11) | −0.0313 (2) | 0.0347 (4) | |
| H14A | −0.1437 | 0.8768 | −0.0996 | 0.042* | |
| C15 | 0.02401 (12) | 0.89096 (10) | −0.1047 (2) | 0.0287 (4) | |
| C16 | 0.01405 (13) | 0.92803 (11) | −0.2857 (2) | 0.0312 (4) | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| N1 | 0.0267 (7) | 0.0273 (7) | 0.0341 (8) | 0.0004 (5) | −0.0016 (6) | 0.0020 (6) |
| O1 | 0.0419 (7) | 0.0307 (7) | 0.0547 (8) | 0.0014 (5) | 0.0029 (6) | −0.0026 (6) |
| O2 | 0.0366 (7) | 0.0296 (7) | 0.0460 (7) | 0.0010 (5) | −0.0007 (5) | −0.0008 (5) |
| O3 | 0.0290 (6) | 0.0499 (8) | 0.0340 (7) | 0.0018 (5) | 0.0007 (5) | 0.0038 (5) |
| O4 | 0.0280 (6) | 0.0490 (8) | 0.0440 (8) | −0.0021 (5) | −0.0058 (5) | 0.0071 (6) |
| C1 | 0.0243 (8) | 0.0398 (10) | 0.0272 (8) | 0.0010 (7) | 0.0044 (6) | 0.0034 (7) |
| C2 | 0.0275 (9) | 0.0479 (11) | 0.0336 (9) | −0.0039 (7) | 0.0043 (7) | −0.0020 (8) |
| C3 | 0.0274 (9) | 0.0706 (14) | 0.0323 (10) | −0.0020 (8) | 0.0019 (7) | 0.0002 (9) |
| C4 | 0.0269 (9) | 0.0720 (14) | 0.0384 (11) | 0.0109 (8) | 0.0020 (8) | 0.0151 (10) |
| C5 | 0.0332 (9) | 0.0469 (11) | 0.0423 (10) | 0.0091 (8) | 0.0091 (8) | 0.0115 (8) |
| C6 | 0.0263 (9) | 0.0392 (10) | 0.0306 (9) | 0.0026 (7) | 0.0063 (7) | 0.0070 (7) |
| C7 | 0.0325 (9) | 0.0307 (10) | 0.0348 (10) | 0.0033 (7) | 0.0078 (7) | 0.0039 (7) |
| C8 | 0.0276 (8) | 0.0323 (9) | 0.0287 (9) | −0.0020 (7) | 0.0046 (7) | 0.0010 (7) |
| C9 | 0.0279 (9) | 0.0318 (9) | 0.0313 (9) | −0.0011 (6) | −0.0019 (7) | 0.0012 (7) |
| C10 | 0.0303 (8) | 0.0202 (8) | 0.0333 (9) | −0.0005 (6) | 0.0029 (7) | −0.0028 (6) |
| C11 | 0.0365 (9) | 0.0283 (9) | 0.0352 (10) | 0.0004 (7) | 0.0014 (7) | −0.0013 (7) |
| C12 | 0.0458 (10) | 0.0276 (9) | 0.0370 (10) | −0.0019 (7) | 0.0099 (8) | 0.0004 (7) |
| C13 | 0.0375 (10) | 0.0310 (9) | 0.0483 (11) | −0.0036 (7) | 0.0138 (8) | −0.0006 (8) |
| C14 | 0.0300 (9) | 0.0287 (9) | 0.0450 (11) | 0.0004 (7) | 0.0027 (7) | −0.0037 (7) |
| C15 | 0.0285 (8) | 0.0218 (8) | 0.0354 (9) | −0.0003 (6) | 0.0015 (7) | −0.0039 (7) |
| C16 | 0.0282 (9) | 0.0279 (9) | 0.0364 (9) | 0.0015 (7) | −0.0013 (7) | −0.0037 (7) |
Geometric parameters (Å, º) top
| O1—C7 | 1.2101 (18) | C6—C7 | 1.492 (2) |
| O2—C8 | 1.2153 (17) | C1—C6 | 1.384 (2) |
| O3—C16 | 1.2279 (17) | C1—C8 | 1.4829 (19) |
| O3—O4i | 2.6361 (16) | N1—C8 | 1.3965 (19) |
| O4—C16 | 1.3224 (17) | C9—H9A | 0.9900 |
| O4—H4B | 0.95 (2) | C9—H9B | 0.9900 |
| C1—C2 | 1.379 (2) | C10—C11 | 1.394 (2) |
| C2—C3 | 1.388 (2) | C10—C15 | 1.402 (2) |
| C2—H2A | 0.9500 | C11—C12 | 1.385 (2) |
| C3—C4 | 1.387 (3) | C11—H11A | 0.9500 |
| C3—H3A | 0.9500 | C12—C13 | 1.379 (2) |
| C4—C5 | 1.392 (2) | C12—H12A | 0.9500 |
| C4—H4A | 0.9500 | C13—C14 | 1.379 (2) |
| C5—C6 | 1.382 (2) | C13—H13A | 0.9500 |
| C5—H5A | 0.9500 | C14—C15 | 1.402 (2) |
| C9—C10 | 1.524 (2) | C14—H14A | 0.9500 |
| N1—C9 | 1.4536 (18) | C15—C16 | 1.480 (2) |
| N1—C7 | 1.3976 (19) | | |
| | | |
| C8—N1—C7 | 111.97 (12) | N1—C8—C1 | 106.01 (13) |
| C8—N1—C9 | 122.74 (12) | N1—C9—C10 | 114.30 (13) |
| C7—N1—C9 | 125.07 (12) | N1—C9—H9A | 108.7 |
| C16—O3—O4i | 126.80 (10) | C10—C9—H9A | 108.7 |
| C16—O4—H4B | 108.9 (12) | N1—C9—H9B | 108.7 |
| C2—C1—C6 | 121.91 (14) | C10—C9—H9B | 108.7 |
| C2—C1—C8 | 129.85 (15) | H9A—C9—H9B | 107.6 |
| C1—C2—C3 | 117.25 (16) | C11—C10—C15 | 118.13 (14) |
| C1—C2—H2A | 121.4 | C11—C10—C9 | 120.28 (13) |
| C3—C2—H2A | 121.4 | C15—C10—C9 | 121.59 (14) |
| C4—C3—C2 | 120.85 (16) | C12—C11—C10 | 121.38 (14) |
| C4—C3—H3A | 119.6 | C12—C11—H11A | 119.3 |
| C2—C3—H3A | 119.6 | C10—C11—H11A | 119.3 |
| C3—C4—C5 | 121.80 (15) | C13—C12—C11 | 120.09 (16) |
| C3—C4—H4A | 119.1 | C13—C12—H12A | 120.0 |
| C5—C4—H4A | 119.1 | C11—C12—H12A | 120.0 |
| C6—C5—C4 | 116.81 (17) | C14—C13—C12 | 119.94 (15) |
| C6—C5—H5A | 121.6 | C14—C13—H13A | 120.0 |
| C4—C5—H5A | 121.6 | C12—C13—H13A | 120.0 |
| C5—C6—C1 | 121.38 (15) | C13—C14—C15 | 120.39 (14) |
| C5—C6—C7 | 130.35 (16) | C13—C14—H14A | 119.8 |
| O1—C7—N1 | 124.23 (14) | C15—C14—H14A | 119.8 |
| O1—C7—C6 | 130.25 (14) | C10—C15—C14 | 120.05 (14) |
| N1—C7—C6 | 105.52 (13) | C10—C15—C16 | 121.64 (14) |
| C1—C6—C7 | 108.25 (12) | C14—C15—C16 | 118.25 (13) |
| C6—C1—C8 | 108.24 (13) | O3—C16—O4 | 121.61 (15) |
| O2—C8—N1 | 124.49 (14) | O3—C16—C15 | 123.59 (14) |
| O2—C8—C1 | 129.50 (14) | O4—C16—C15 | 114.78 (14) |
| | | |
| C11—C10—C9—N1 | −5.3 (2) | C10—C9—N1—C7 | 86.64 (17) |
| Symmetry code: (i) −x, −y+2, −z−1. |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| O4—H4B···O3i | 0.95 (2) | 1.69 (2) | 2.6361 (16) | 171 (2) |
| Symmetry code: (i) −x, −y+2, −z−1. |
Experimental details
| | (I) | (II) | (III) |
| Crystal data |
| Chemical formula | C14H10N4 | C14H14O8 | C16H11NO4 |
| Mr | 234.26 | 310.25 | 281.26 |
| Crystal system, space group | Monoclinic, P21/c | Monoclinic, P21/n | Monoclinic, P21/c |
| Temperature (K) | 153 | 153 | 153 |
| a, b, c (Å) | 8.818 (1), 8.377 (1), 8.174 (1) | 9.2233 (7), 11.747 (1), 13.936 (1) | 12.125 (1), 13.720 (1), 7.7247 (8) |
| β (°) | 108.51 (1) | 90.451 (1) | 96.722 (2) |
| V (Å3) | 572.48 (15) | 1509.9 (2) | 1276.2 (2) |
| Z | 2 | 4 | 4 |
| Radiation type | Mo Kα | Mo Kα | Mo Kα |
| µ (mm−1) | 0.09 | 0.11 | 0.11 |
| Crystal size (mm) | 0.12 × 0.11 × 0.06 | 0.41 × 0.30 × 0.27 | 0.33 × 0.17 × 0.08 |
| |
| Data collection |
| Diffractometer | Bruker SMART 1000 CCD
diffractometer | Bruker SMART 1000 CCD
diffractometer | Bruker Smart 1000 CCD
diffractometer |
| Absorption correction | Numerical
face-indexed (SHELXTL/PC; Sheldrick, 1997) | Numerical
face-indexed (SHELXTL/PC; Sheldrick, 1997) | Numerical
face indexed (Sheldrick, 1999) |
| Tmin, Tmax | 0.985, 0.995 | 0.959, 0.976 | 0.974, 0.992 |
No. of measured, independent and
observed [I > 2σ(I)] reflections | 3540, 1335, 836 | 8088, 2789, 2320 | 6929, 2365, 1722 |
| Rint | 0.035 | 0.016 | 0.026 |
| (sin θ/λ)max (Å−1) | 0.665 | 0.606 | 0.606 |
| |
| Refinement |
| R[F2 > 2σ(F2)], wR(F2), S | 0.044, 0.098, 1.00 | 0.043, 0.116, 1.84 | 0.036, 0.089, 1.10 |
| No. of reflections | 1335 | 2789 | 2365 |
| No. of parameters | 82 | 203 | 194 |
| H-atom treatment | H-atom parameters constrained | H-atom parameters constrained | H atoms; see below |
| Δρmax, Δρmin (e Å−3) | 0.26, −0.16 | 0.27, −0.26 | 0.16, −0.17 |
Selected geometric parameters (Å, º) for (I) top| C1—C2 | 1.397 (2) | C4—C5 | 1.464 (2) |
| C2—C3 | 1.396 (2) | C6—C7 | 1.459 (2) |
| C3—C1i | 1.390 (2) | C5—N1 | 1.149 (2) |
| C1—C4 | 1.518 (2) | C7—N2 | 1.1447 (19) |
| C2—C6 | 1.522 (2) | | |
| | | |
| C3i—C1—C2 | 119.42 (13) | C3—C2—C6 | 121.05 (14) |
| C3—C2—C1 | 118.44 (14) | N1—C5—C4 | 179.67 (17) |
| C1i—C3—C2 | 122.14 (14) | N2—C7—C6 | 179.8 (2) |
| | | |
| C2—C1—C4—C5 | −67.20 (19) | C1—C4—C5—N1 | 16 (34) |
| Symmetry code: (i) −x+1, −y+1, −z. |
Selected geometric parameters (Å, º) for (II) top| C1—C2 | 1.395 (2) | C11—O6 | 1.186 (2) |
| C2—C3 | 1.370 (2) | C13—O8 | 1.1918 (19) |
| C3—C4 | 1.387 (2) | C3—O3 | 1.4006 (17) |
| C4—C5 | 1.383 (2) | C4—O5 | 1.3939 (17) |
| C5—C6 | 1.386 (2) | C5—O7 | 1.3937 (17) |
| C1—C6 | 1.390 (2) | C9—O3 | 1.3751 (18) |
| C1—C7 | 1.492 (2) | C11—O5 | 1.3646 (18) |
| C7—O2 | 1.2061 (18) | C13—O7 | 1.3719 (19) |
| C9—O4 | 1.1910 (19) | | |
| | | |
| O2—C7—O1 | 123.63 (14) | C7—O1—C8 | 115.96 (13) |
| O4—C9—C10 | 127.43 (15) | C9—O3—C3 | 115.95 (11) |
| O6—C11—C12 | 127.67 (15) | C11—O5—C4 | 116.60 (11) |
| O8—C13—C14 | 126.61 (15) | C13—O7—C5 | 119.24 (12) |
| | | |
| C2—C3—O3—C9 | 83.22 (17) | C4—C5—O7—C13 | −133.03 (14) |
| C3—C4—O5—C11 | −84.37 (17) | C6—C1—C7—O1 | −3.2 (2) |
Selected geometric parameters (Å, º) for (III) top| O1—C7 | 1.2101 (18) | N1—C7 | 1.3976 (19) |
| O2—C8 | 1.2153 (17) | C6—C7 | 1.492 (2) |
| O3—C16 | 1.2279 (17) | C1—C6 | 1.384 (2) |
| O4—C16 | 1.3224 (17) | C1—C8 | 1.4829 (19) |
| C9—C10 | 1.524 (2) | N1—C8 | 1.3965 (19) |
| N1—C9 | 1.4536 (18) | | |
| | | |
| C8—N1—C7 | 111.97 (12) | N1—C8—C1 | 106.01 (13) |
| N1—C7—C6 | 105.52 (13) | N1—C9—C10 | 114.30 (13) |
| C1—C6—C7 | 108.25 (12) | O3—C16—O4 | 121.61 (15) |
| C6—C1—C8 | 108.24 (13) | | |
| | | |
| C10—C9—N1—C7 | 86.64 (17) | | |
Hydrogen-bond geometry (Å, º) for (III) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| O4—H4B···O3i | 0.95 (2) | 1.69 (2) | 2.6361 (16) | 171 (2) |
| Symmetry code: (i) −x, −y+2, −z−1. |
 O contact is formed, with an H
O contact is formed, with an H O distance of 2.34 Å. The structure of 2-(N-phthalimidomethyl)benzoic acid, C16H11NO4, reveals hydrogen-bonded dimers linked by the carboxyl groups of adjacent molecules. The O4
O distance of 2.34 Å. The structure of 2-(N-phthalimidomethyl)benzoic acid, C16H11NO4, reveals hydrogen-bonded dimers linked by the carboxyl groups of adjacent molecules. The O4 O3 distance is 2.636 (2) Å and the O4—H
O3 distance is 2.636 (2) Å and the O4—H O3 angle is 171 (2)°.
O3 angle is 171 (2)°.
 journal menu
journal menu








![[Figure 1]](http://journals.iucr.org/c/issues/2000/02/00/bk1493/bk1493fig1thm.gif)
![[Figure 2]](http://journals.iucr.org/c/issues/2000/02/00/bk1493/bk1493fig2thm.gif)
![[Figure 3]](http://journals.iucr.org/c/issues/2000/02/00/bk1493/bk1493fig3thm.gif)
![[Figure 4]](http://journals.iucr.org/c/issues/2000/02/00/bk1493/bk1493fig4thm.gif)




Derivatized porphyrins play useful roles as models for protein active sites. Particular effort has been paid to models for the heme active sites of myoglobin and hemoglobin (Momenteau & Reed, 1994). This laboratory has synthesized and characterized a number of four-atom-linked capped porphyrins (Johnson et al., 1996), five-atom-linked capped porphyrins (Ma et al., 1996), and five-plus-atom-linked capped porphyrins (Slebodnick et al., 1996). Herein, we report the structures of three precursors characterized in the course of this research, 1,2,4,5-tetrakis(cyanomethyl)benzene, (I), methyl 3,4,5-triacetoxybenzoate, (II) and 2-(N-phthalimidomethyl)benzoic acid, (III). \scheme
Compound (I) was prepared as a precursor for the synthesis of a three-atom-linked capped porphyrin. Fig. 1 is a displacement ellipsoid diagram of (I). The molecule sits on an inversion centre and its symmetry-generated ellipsoids are shown without labels. The benzene ring is planar by symmetry. Selected bond lengths and angles are listed in Table 1. One of the unique cyanomethyl groups remains nearly in the plane of the benzene ring, and this cyanomethyl group is linear, with a C6—C7—N2 angle of 179.8 (2) Å. The other unique cyanomethyl group is twisted 67.2 (2)° relative to the C1—C2 bond in the benzene ring (Table 1). This cyanomethyl group is also linear, with a bond angle of 179.67 (17) Å for C4—C5—N1. Viewed down the c axis, molecules of (I) form long straight columns aligned along the centroids of the benzene rings. The molecules in each column pack tilted at 72.8 (1)° relative to molecules in adjacent columns.
The structure of 1,2,4,5-tetramethylbenzene, durene, has been examined both as a donor in charge transfer complexes (Lefebvre, 1989) and to understand equilibrium conformations of the molecule (Prince et al., 1973, and references therein). There has been interest in 1,2,4,5-tetracyanobenzene (TCNB) as an acceptor in charge transfer complexes (Lefebvre, 1989, and references therein). In the series durene, (I) and TCNB, there is a shift from mildly electron-donating methyl substituents to strongly electron-withdrawing cyano substituents. The changes in electron donation and conjugation have no significant effect on the C≐C bond lengths in the benzene ring. However, durene structures and (I) exhibit geometric distortion of the benzene ring, presumably resulting from the bulky substituents. In durene structures the angle formed by the unsubstituted C atoms with their nearest C neighbours is approximately 122–123°, with a corresponding reduction of other ring angles. For (I), this angle is 122.14 (14)°.
Compound (II) (Fig. 2) was prepared as a precursor for a capped porphyrin with asymmetric linkages between the benzene `cap' and the porphyrin. In (II), the central benzene ring is planar, with a maximum deviation from the mean plane of 0.007 (1) Å for atom C5. Selected bond lengths and angles are listed in Table 2. The three acetoxy groups are rotated out of the benzene plane by 83.22 (17), −84.37 (17) and −133.03 (14)°, respectively (as measured by torsion angles). In contrast, the carbomethoxy group remains nearly in the plane, with a torsion angle of −3.2 (2)°.
Molecules of (II) are associated in two ways. The first involves a close contact between O2Ai···H10A—C10 [symmetry code: (i) x − 1/2, −1/2 − y, z − 1/2]. The C10—H10A distance is 0.98 Å (fixed with a riding model, see refinement details below) and the O2Ai···H10A distance is 2.34 Å, while the O2Ai···H10A—C10 bond angle is 170°. Molecules of (II) form chains oriented along these contacts and molecules in the chain tilt back and forth 89.6 (1)° relative to each other. The second association is dimeric: molecules generated by (x, y, z) and (-x, 2 − y, 1 − z) form offset dimers. The benzene rings are parallel, but the molecules are rotated 180° with respect to each other and the benzene centroids are slightly offset. The separation between the benzene planes is 3.29 (1) Å.
Compound (III) (Fig. 3) was prepared as a precursor for capped porphyrins containing N atoms at an intermediate position on the `arms' linking the benzene `cap' to the porphyrin. Compound (III) is a primary amine of the form RNH2 protected with phthalimide. The phthalimide group is planar, with a maximum deviation from the 11-atom mean plane of 0.013 (1) Å for atom C6. The benzene of the benzyl group is also planar, with a maximum deviation of 0.007 (1) Å for atom C12. The dihedral angle between these planes is 80.86 (4)°. Selected bond lengths and angles are listed in Table 3. The twist in the molecule occurs at atom C9: the C10—C9—N1—C7 torsion angle is 86.6 (2)°. Molecules of (III) form hydrogen-bonded dimers through their carboxyl groups. The O4—H4B distance is 0.90 (3) Å and the H4B···O3Aii distance is 1.77 (3) Å [symmetry code: (ii) −x, −y + 2,-z − 1]. The O4···O3Aii distance is 2.638 (2) Å, and the O4—H4B···O3Aii bond angle is 163 (2)°. This short distance is normal for strongly hydrogen-bonded carboxylic acid dimers in the crystalline state (Speakman, 1972). Molecules of (III) pack with alternating layers of hydrogen-bonded carboxylbenzyl groups and stacked phthalimide groups (Fig. 4).