In the title compound, [CuCl
2(C
3H
5N
7)
2], the coordination polyhedron of the Cu atom is an elongated square bipyramid with
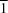
site symmetry. The equatorial positions are occupied by the two Cl atoms with Cu—Cl distances of 2.288 (1) Å and two azidoethyltetrazole ligands with Cu—N distances of 1.999 (2) Å. Two Cl atoms in axial positions are 2.956 (1) Å distant from the Cu atom. The Cl atoms play the role of non-symmetrical bridges responsible for formation of layers parallel to the
bc plane.
Supporting information
CCDC reference: 170862
Key indicators
- Single-crystal X-ray study
- T = 293 K
- Mean
![[sigma]](//journals.iucr.org/logos/entities/sigma_rmgif.gif) (C-C) = 0.004 Å
(C-C) = 0.004 Å
- R factor = 0.033
- wR factor = 0.097
- Data-to-parameter ratio = 16.4
checkCIF results
No syntax errors found
ADDSYM reports no extra symmetry
 Alert Level C:
Alert Level C:
PLAT_731 Alert C Bond Calc 2.956(3), Rep 2.9560(10) .... 3.00 s.u-Ratio
CU1 -CL1 1.555 2.545
0 Alert Level A = Potentially serious problem
0 Alert Level B = Potential problem
1 Alert Level C = Please check
The synthesis of the title compound was carried out at 425–426 K by reaction of
CuCl2·2H2O with 1-(2-azidoethyl)tetrazole (molar ratio 1:2) by a
method proposed by Degtiarik et al. (1985). Recrystallization of (I)
was performed from water solution. 1-(2-Azidoethyl)tetrazole was prepared by
heterocyclization of 1-amino-2-azidoethan with ethyl orthoformate and sodium
azide in acetic acid (Gaponik et al., 1985).
The H– toms were included in geometrically calculated positions and refined
using a riding model with Uiso(H) = 1.2Ueq of the
corresponding carrier atom.
Data collection: R3m Software (Nicolet, 1980); cell refinement: R3m software; data reduction: R3m software; program(s) used to solve structure: SIR97 (Altomare et al., 1999); program(s) used to refine structure: SHELXL97 (Sheldrick, 1997); molecular graphics: ORTEP-3 for Windows (Farrugia, 1997); software used to prepare material for publication: SHELXL97.
catena-poly[[bis(1-(2-azidoethyl)tetrazole-
N4)-copper(II)]-di-µ-chloro]
top
Crystal data top
| [CuCl2(C3H5N7)2] | F(000) = 414 |
| Mr = 412.72 | Dx = 1.818 Mg m−3 |
| Monoclinic, P21/c | Mo Kα radiation, λ = 0.71073 Å |
| a = 15.616 (2) Å | Cell parameters from 25 reflections |
| b = 6.702 (2) Å | θ = 17.2–24.2° |
| c = 7.208 (6) Å | µ = 1.83 mm−1 |
| β = 91.87 (3)° | T = 293 K |
| V = 754.0 (7) Å3 | Plate, blue |
| Z = 2 | 0.80 × 0.46 × 0.02 mm |
Data collection top
Nicolet R3m four-circle
diffractometer | 1529 reflections with I > 2σ(I) |
| Radiation source: fine-focus sealed tube | Rint = 0.018 |
| Graphite monochromator | θmax = 27.5°, θmin = 1.3° |
| ω/2θ scans | h = −20→20 |
Absorption correction: ψ scan
(North et al., 1968) | k = −8→0 |
| Tmin = 0.323, Tmax = 0.964 | l = 0→9 |
| 1880 measured reflections | 3 standard reflections every 100 reflections |
| 1734 independent reflections | intensity decay: none |
Refinement top
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.033 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.097 | H-atom parameters constrained |
| S = 1.04 | w = 1/[σ2(Fo2) + (0.069P)2 + 0.223P]
where P = (Fo2 + 2Fc2)/3 |
| 1734 reflections | (Δ/σ)max < 0.001 |
| 106 parameters | Δρmax = 0.47 e Å−3 |
| 0 restraints | Δρmin = −0.34 e Å−3 |
Crystal data top
| [CuCl2(C3H5N7)2] | V = 754.0 (7) Å3 |
| Mr = 412.72 | Z = 2 |
| Monoclinic, P21/c | Mo Kα radiation |
| a = 15.616 (2) Å | µ = 1.83 mm−1 |
| b = 6.702 (2) Å | T = 293 K |
| c = 7.208 (6) Å | 0.80 × 0.46 × 0.02 mm |
| β = 91.87 (3)° | |
Data collection top
Nicolet R3m four-circle
diffractometer | 1529 reflections with I > 2σ(I) |
Absorption correction: ψ scan
(North et al., 1968) | Rint = 0.018 |
| Tmin = 0.323, Tmax = 0.964 | 3 standard reflections every 100 reflections |
| 1880 measured reflections | intensity decay: none |
| 1734 independent reflections | |
Refinement top
| R[F2 > 2σ(F2)] = 0.033 | 0 restraints |
| wR(F2) = 0.097 | H-atom parameters constrained |
| S = 1.04 | Δρmax = 0.47 e Å−3 |
| 1734 reflections | Δρmin = −0.34 e Å−3 |
| 106 parameters | |
Special details top
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes)
are estimated using the full covariance matrix. The cell e.s.d.'s are taken
into account individually in the estimation of e.s.d.'s in distances, angles
and torsion angles; correlations between e.s.d.'s in cell parameters are only
used when they are defined by crystal symmetry. An approximate (isotropic)
treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s.
planes. |
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor
wR and goodness of fit S are based on F2, conventional
R-factors R are based on F, with F set to zero for
negative F2. The threshold expression of F2 >
σ(F2) is used only for calculating R-factors(gt) etc.
and is not relevant to the choice of reflections for refinement.
R-factors based on F2 are statistically about twice as large
as those based on F, and R- factors based on ALL data will be
even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| Cu1 | 0.0000 | 0.5000 | 0.5000 | 0.02654 (15) | |
| Cl1 | −0.05746 (3) | 0.71706 (7) | 0.28493 (7) | 0.02841 (16) | |
| N1 | 0.20604 (11) | 0.8732 (3) | 0.4690 (3) | 0.0315 (4) | |
| N2 | 0.24478 (14) | 0.6972 (3) | 0.4424 (4) | 0.0463 (6) | |
| N3 | 0.18738 (13) | 0.5609 (3) | 0.4586 (4) | 0.0427 (5) | |
| N4 | 0.11123 (10) | 0.6469 (3) | 0.4928 (2) | 0.0259 (4) | |
| C5 | 0.12436 (13) | 0.8398 (3) | 0.4984 (3) | 0.0301 (4) | |
| H5 | 0.0832 | 0.9367 | 0.5196 | 0.036* | |
| C6 | 0.25272 (17) | 1.0620 (4) | 0.4532 (5) | 0.0462 (6) | |
| H6A | 0.2165 | 1.1709 | 0.4920 | 0.055* | |
| H6B | 0.2664 | 1.0841 | 0.3245 | 0.055* | |
| C7 | 0.33252 (18) | 1.0609 (5) | 0.5685 (5) | 0.0493 (7) | |
| H7A | 0.3692 | 0.9516 | 0.5327 | 0.059* | |
| H7B | 0.3198 | 1.0459 | 0.6986 | 0.059* | |
| N5 | 0.37491 (16) | 1.2541 (4) | 0.5364 (5) | 0.0634 (8) | |
| N6 | 0.44506 (17) | 1.2736 (4) | 0.6084 (4) | 0.0605 (7) | |
| N7 | 0.5109 (2) | 1.3094 (7) | 0.6666 (6) | 0.1060 (14) | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| Cu1 | 0.0247 (2) | 0.0269 (2) | 0.0280 (2) | −0.00358 (12) | 0.00057 (14) | 0.00865 (13) |
| Cl1 | 0.0346 (3) | 0.0283 (3) | 0.0223 (3) | 0.00035 (17) | 0.00029 (19) | 0.00461 (18) |
| N1 | 0.0296 (9) | 0.0286 (9) | 0.0364 (11) | −0.0046 (7) | 0.0016 (7) | 0.0013 (7) |
| N2 | 0.0326 (10) | 0.0355 (10) | 0.0718 (17) | −0.0034 (8) | 0.0140 (10) | −0.0078 (10) |
| N3 | 0.0303 (9) | 0.0305 (9) | 0.0679 (16) | −0.0010 (8) | 0.0093 (9) | −0.0085 (10) |
| N4 | 0.0263 (8) | 0.0268 (8) | 0.0248 (9) | −0.0013 (6) | 0.0034 (6) | 0.0033 (7) |
| C5 | 0.0285 (10) | 0.0282 (9) | 0.0337 (12) | −0.0016 (8) | 0.0008 (8) | 0.0018 (8) |
| C6 | 0.0394 (13) | 0.0352 (12) | 0.0635 (18) | −0.0136 (11) | −0.0071 (12) | 0.0120 (12) |
| C7 | 0.0439 (14) | 0.0448 (13) | 0.0585 (18) | −0.0159 (12) | −0.0088 (13) | 0.0069 (13) |
| N5 | 0.0509 (15) | 0.0575 (14) | 0.081 (2) | −0.0275 (12) | −0.0146 (14) | 0.0163 (13) |
| N6 | 0.0510 (14) | 0.0704 (17) | 0.0600 (17) | −0.0302 (13) | 0.0016 (12) | 0.0021 (13) |
| N7 | 0.064 (2) | 0.141 (4) | 0.111 (3) | −0.052 (2) | −0.024 (2) | 0.010 (3) |
Geometric parameters (Å, º) top
| Cu1—Cl1 | 2.288 (1) | C5—H5 | 0.9300 |
| Cu1—Cl1i | 2.956 (1) | C6—C7 | 1.475 (4) |
| Cu1—N4 | 1.999 (2) | C6—H6A | 0.9700 |
| N1—N2 | 1.342 (3) | C6—H6B | 0.9700 |
| N1—C5 | 1.319 (3) | C7—N5 | 1.476 (3) |
| N1—C6 | 1.467 (3) | C7—H7A | 0.9700 |
| N2—N3 | 1.288 (3) | C7—H7B | 0.9700 |
| N3—N4 | 1.351 (3) | N5—N6 | 1.204 (4) |
| N4—C5 | 1.309 (3) | N6—N7 | 1.123 (4) |
| | | |
| N4—Cu1—Cl1 | 89.49 (5) | N1—C6—C7 | 111.4 (2) |
| N4ii—Cu1—Cl1i | 89.18 (6) | N1—C6—H6A | 109.3 |
| Cl1—Cu1—Cl1i | 93.35 (5) | C7—C6—H6A | 109.3 |
| C5—N1—N2 | 108.5 (2) | N1—C6—H6B | 109.3 |
| C5—N1—C6 | 130.1 (2) | C7—C6—H6B | 109.3 |
| N2—N1—C6 | 121.3 (2) | H6A—C6—H6B | 108.0 |
| N3—N2—N1 | 107.0 (2) | C6—C7—N5 | 106.4 (2) |
| N2—N3—N4 | 109.5 (2) | C6—C7—H7A | 110.4 |
| C5—N4—N3 | 106.8 (2) | N5—C7—H7A | 110.4 |
| C5—N4—Cu1 | 128.4 (1) | C6—C7—H7B | 110.4 |
| N3—N4—Cu1 | 124.4 (1) | N5—C7—H7B | 110.4 |
| N4—C5—N1 | 108.3 (2) | H7A—C7—H7B | 108.6 |
| N4—C5—H5 | 125.9 | N6—N5—C7 | 115.7 (3) |
| N1—C5—H5 | 125.9 | N7—N6—N5 | 173.1 (4) |
| | | |
| C5—N1—N2—N3 | −0.9 (3) | Cl1i—Cu1—N4—N3 | −32.29 (18) |
| C6—N1—N2—N3 | −177.7 (2) | N3—N4—C5—N1 | −0.1 (3) |
| N1—N2—N3—N4 | 0.9 (3) | Cu1—N4—C5—N1 | −172.95 (15) |
| N2—N3—N4—C5 | −0.5 (3) | N2—N1—C5—N4 | 0.6 (3) |
| N2—N3—N4—Cu1 | 172.71 (18) | C6—N1—C5—N4 | 177.0 (2) |
| Cl1—Cu1—N4—C5 | 46.07 (19) | C5—N1—C6—C7 | 132.1 (3) |
| Cl1ii—Cu1—N4—C5 | −133.93 (19) | N2—N1—C6—C7 | −51.9 (4) |
| Cl1i—Cu1—N4—C5 | 139.41 (18) | N1—C6—C7—N5 | 178.1 (3) |
| Cl1—Cu1—N4—N3 | −125.63 (19) | C6—C7—N5—N6 | −174.3 (3) |
| Cl1ii—Cu1—N4—N3 | 54.37 (19) | | |
| Symmetry codes: (i) −x, y−1/2, −z+1/2; (ii) −x, −y+1, −z+1. |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| C5—H5···Cl1iii | 0.93 | 2.75 | 3.529 (3) | 142 |
| Symmetry code: (iii) −x, −y+2, −z+1. |
Experimental details
| Crystal data |
| Chemical formula | [CuCl2(C3H5N7)2] |
| Mr | 412.72 |
| Crystal system, space group | Monoclinic, P21/c |
| Temperature (K) | 293 |
| a, b, c (Å) | 15.616 (2), 6.702 (2), 7.208 (6) |
| β (°) | 91.87 (3) |
| V (Å3) | 754.0 (7) |
| Z | 2 |
| Radiation type | Mo Kα |
| µ (mm−1) | 1.83 |
| Crystal size (mm) | 0.80 × 0.46 × 0.02 |
| |
| Data collection |
| Diffractometer | Nicolet R3m four-circle
diffractometer |
| Absorption correction | ψ scan
(North et al., 1968) |
| Tmin, Tmax | 0.323, 0.964 |
No. of measured, independent and
observed [I > 2σ(I)] reflections | 1880, 1734, 1529 |
| Rint | 0.018 |
| (sin θ/λ)max (Å−1) | 0.650 |
| |
| Refinement |
| R[F2 > 2σ(F2)], wR(F2), S | 0.033, 0.097, 1.04 |
| No. of reflections | 1734 |
| No. of parameters | 106 |
| H-atom treatment | H-atom parameters constrained |
| Δρmax, Δρmin (e Å−3) | 0.47, −0.34 |
Selected geometric parameters (Å, º) top| Cu1—Cl1 | 2.288 (1) | N3—N4 | 1.351 (3) |
| Cu1—Cl1i | 2.956 (1) | N4—C5 | 1.309 (3) |
| Cu1—N4 | 1.999 (2) | C6—C7 | 1.475 (4) |
| N1—N2 | 1.342 (3) | C7—N5 | 1.476 (3) |
| N1—C5 | 1.319 (3) | N5—N6 | 1.204 (4) |
| N1—C6 | 1.467 (3) | N6—N7 | 1.123 (4) |
| N2—N3 | 1.288 (3) | | |
| | | |
| N4—Cu1—Cl1 | 89.49 (5) | C5—N4—N3 | 106.8 (2) |
| N4ii—Cu1—Cl1i | 89.18 (6) | C5—N4—Cu1 | 128.4 (1) |
| Cl1—Cu1—Cl1i | 93.35 (5) | N3—N4—Cu1 | 124.4 (1) |
| C5—N1—N2 | 108.5 (2) | N4—C5—N1 | 108.3 (2) |
| C5—N1—C6 | 130.1 (2) | N1—C6—C7 | 111.4 (2) |
| N2—N1—C6 | 121.3 (2) | C6—C7—N5 | 106.4 (2) |
| N3—N2—N1 | 107.0 (2) | N6—N5—C7 | 115.7 (3) |
| N2—N3—N4 | 109.5 (2) | N7—N6—N5 | 173.1 (4) |
| Symmetry codes: (i) −x, y−1/2, −z+1/2; (ii) −x, −y+1, −z+1. |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| C5—H5···Cl1iii | 0.93 | 2.75 | 3.529 (3) | 141.8 |
| Symmetry code: (iii) −x, −y+2, −z+1. |

 journal menu
journal menu











![[Figure 1]](http://journals.iucr.org/e/issues/2001/08/00/ya6036/ya6036fig1thm.gif)
![[Figure 2]](http://journals.iucr.org/e/issues/2001/08/00/ya6036/ya6036fig2thm.gif)




It has been found in the past few years that the complexes of copper(II) chloride with 1-monosubstituted tetrazole of composition CuCl2L2, where L = 1-ethyl-, 1-hexyl, 1-vinyl and 1-allyltetrazole, undergo magnetic phase transition to ferromagnetic form at T = 10–12 K (Gaponik, 1998). X-ray investigations of the complexes with L = 1-ethyltetrazole (Virovets et al., 1995) and L = 1-allyltetrazole (Virovets et al., 1996) showed layered polymeric crystal structure of these compounds, which is known to be a necessary condition for ferromagnetic ordering for transition metal complexes (Ovcharenko & Sagdeev, 1999). In the present work, the crystal structure of a new complex, (I), of composition CuCl2L2 (L = 1-azidoethyltetrazole), is reported.
In the title compound, the coordination polyhedron of the Cu atom is an elongated square bipyramid with 1 site symmetry. The equatorial positions are occupied by the two Cl atoms with Cu—Cl distances of 2.288 (1) Å and two N4 atoms of azidoethyltetrazole molecules with Cu—N distances of 1.999 (2) Å. Two Cl atoms in axial position are at a distance of 2.956 (1) Å from the Cu atom.
The tetrazole ring is essentially planar, with a mean deviation of terazole ring atoms from their least-squares plane of 0.006 (2) Å. The tetrazole plane forms the dihedral angle of 50.9 (1)° with the equatorial plane of the coordination square pyramid of the Cu atom. In the azide fragment, the N5—N6 distance is 1.204 (4) Å and the N6—N7 distance is 1.123 (4) Å; the N5—N6—N7 and C7—N5—N6 bond angles are 173.1 (4) and 115.7 (3)°, respectively.
The analysis of the crystal packing of complex (I) reveals a couple of interesting features. The Cl atoms play the role of non-symmetrical bridges which are responsible for formation of layers parallel to the yz plane. They also act as acceptors of non-classic intermolecular hydrogen bonds C5—H5···Cl, with a Cl···C5 distance of 3.529 (3) Å (Steiner, 1996). Thus, the title compound has a layered polymeric crystal structure, and hence may be an object for magnetic properties investigations.