Related literature
For a related structure, see: Khan et al. (2011![Khan, I. U., Sheikh, T. A., Ejaz & Harrison, W. T. A. (2011). Acta Cryst. E67, o2371. [Khan, I. U., Sheikh, T. A., Ejaz & Harrison, W. T. A. (2011). Acta Cryst. E67, o2371.]](../../../../../../logos/arrows/e_arr.gif) ).
).
Experimental
Crystal data
C20H20N2O4S2 Mr = 416.50 Monoclinic, C 2/c a = 16.4319 (6) Å b = 7.8024 (3) Å c = 16.1217 (6) Å β = 111.543 (2)° V = 1922.54 (12) Å3 Z = 4 Mo Kα radiation μ = 0.31 mm−1 T = 296 K 0.15 × 0.11 × 0.08 mm
|
Data collection
Bruker APEXII CCD diffractometer 8653 measured reflections 2346 independent reflections 1841 reflections with I > 2σ(I) Rint = 0.024
|
| D—H⋯A | D—H | H⋯A | D⋯A | D—H⋯A | | N1—H1⋯O2i | 0.81 (2) | 2.16 (2) | 2.960 (2) | 166 (2) | | C5—H5⋯O1ii | 0.93 | 2.55 | 3.461 (2) | 166 | | C10—H10⋯O1iii | 0.93 | 2.57 | 3.379 (3) | 146 | Symmetry codes: (i) ![[-x+{\script{1\over 2}}, y-{\script{1\over 2}}, -z+{\script{1\over 2}}]](teximages/su2323fi2.gif) ; (ii) -x+1, -y+1, -z+1; (iii) x, y-1, z. ; (ii) -x+1, -y+1, -z+1; (iii) x, y-1, z.
| |
Data collection: APEX2 (Bruker, 2007![Bruker (2007). APEX2 and SAINT. Bruker AXS Inc., Madison, Wisconsin, USA. [Bruker (2007). APEX2 and SAINT. Bruker AXS Inc., Madison, Wisconsin, USA.]](../../../../../../logos/arrows/e_arr.gif) ); cell refinement: SAINT (Bruker, 2007
); cell refinement: SAINT (Bruker, 2007![Bruker (2007). APEX2 and SAINT. Bruker AXS Inc., Madison, Wisconsin, USA. [Bruker (2007). APEX2 and SAINT. Bruker AXS Inc., Madison, Wisconsin, USA.]](../../../../../../logos/arrows/e_arr.gif) ); data reduction: SAINT; program(s) used to solve structure: SHELXS97 (Sheldrick, 2008
); data reduction: SAINT; program(s) used to solve structure: SHELXS97 (Sheldrick, 2008![Sheldrick, G. M. (2008). Acta Cryst. A64, 112-122. [Sheldrick, G. M. (2008). Acta Cryst. A64, 112-122.]](../../../../../../logos/arrows/e_arr.gif) ); program(s) used to refine structure: SHELXL97 (Sheldrick, 2008
); program(s) used to refine structure: SHELXL97 (Sheldrick, 2008![Sheldrick, G. M. (2008). Acta Cryst. A64, 112-122. [Sheldrick, G. M. (2008). Acta Cryst. A64, 112-122.]](../../../../../../logos/arrows/e_arr.gif) ); molecular graphics: ORTEP-3 (Farrugia, 1997
); molecular graphics: ORTEP-3 (Farrugia, 1997![Farrugia, L. J. (1997). J. Appl. Cryst. 30, 565. [Farrugia, L. J. (1997). J. Appl. Cryst. 30, 565.]](../../../../../../logos/arrows/e_arr.gif) ); software used to prepare material for publication: SHELXL97.
); software used to prepare material for publication: SHELXL97.
Supporting information
The m-xylylenediamine (0.132 ml, 1.0 mmol) was mixed with 20 ml distilled water in a 50 ml round bottom flask. Benzene sulfonyl chloride (0.255 ml, 2.0 mmol) was added while maintaining the pH of the reaction mixture at ca. 9.0 using sodium carbonate solution (3%) and the mixture was stirred for five hours. The white precipitated product was filtered, washed, dried and crystallized from methanol to generate colourless blocks of the title compound.
The N-bound H atom was located in a difference map and its position was freely refined with the constraint Uiso(H) = 1.2Ueq(N). The C-bound hydrogen atoms were placed in calculated positions (C—H = 0.93–0.97 Å) and refined as riding atoms with Uiso(H) = 1.2Ueq(C).
Data collection: APEX2 (Bruker, 2007); cell refinement: SAINT (Bruker, 2007); data reduction: SAINT (Bruker, 2007); program(s) used to solve structure: SHELXS97 (Sheldrick, 2008); program(s) used to refine structure: SHELXL97 (Sheldrick, 2008); molecular graphics: ORTEP-3 (Farrugia, 1997); software used to prepare material for publication: SHELXL97 (Sheldrick, 2008).
N,
N'-[1,3-Phenylenebis(methylene)]dibenzenesulfonamide
top Crystal data top | C20H20N2O4S2 | F(000) = 872 |
| Mr = 416.50 | Dx = 1.439 Mg m−3 |
| Monoclinic, C2/c | Mo Kα radiation, λ = 0.71073 Å |
| Hall symbol: -C 2yc | Cell parameters from 2346 reflections |
| a = 16.4319 (6) Å | θ = 2.9–28.3° |
| b = 7.8024 (3) Å | µ = 0.31 mm−1 |
| c = 16.1217 (6) Å | T = 296 K |
| β = 111.543 (2)° | Block, colourless |
| V = 1922.54 (12) Å3 | 0.15 × 0.11 × 0.08 mm |
| Z = 4 | |
Data collection top Bruker APEXII CCD
diffractometer | 1841 reflections with I > 2σ(I) |
| Radiation source: fine-focus sealed tube | Rint = 0.024 |
| Graphite monochromator | θmax = 28.3°, θmin = 2.9° |
| ω scans | h = −21→21 |
| 8653 measured reflections | k = −9→10 |
| 2346 independent reflections | l = −21→21 |
Refinement top | Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.037 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.103 | H atoms treated by a mixture of independent and constrained refinement |
| S = 1.07 | w = 1/[σ2(Fo2) + (0.0466P)2 + 1.1784P]
where P = (Fo2 + 2Fc2)/3 |
| 2346 reflections | (Δ/σ)max = 0.001 |
| 131 parameters | Δρmax = 0.29 e Å−3 |
| 0 restraints | Δρmin = −0.34 e Å−3 |
Crystal data top | C20H20N2O4S2 | V = 1922.54 (12) Å3 |
| Mr = 416.50 | Z = 4 |
| Monoclinic, C2/c | Mo Kα radiation |
| a = 16.4319 (6) Å | µ = 0.31 mm−1 |
| b = 7.8024 (3) Å | T = 296 K |
| c = 16.1217 (6) Å | 0.15 × 0.11 × 0.08 mm |
| β = 111.543 (2)° | |
Data collection top Bruker APEXII CCD
diffractometer | 1841 reflections with I > 2σ(I) |
| 8653 measured reflections | Rint = 0.024 |
| 2346 independent reflections | |
Refinement top | R[F2 > 2σ(F2)] = 0.037 | 0 restraints |
| wR(F2) = 0.103 | H atoms treated by a mixture of independent and constrained refinement |
| S = 1.07 | Δρmax = 0.29 e Å−3 |
| 2346 reflections | Δρmin = −0.34 e Å−3 |
| 131 parameters | |
Special details top Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes. |
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2, conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2 are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| C1 | 0.23478 (13) | 0.4749 (3) | 0.57485 (13) | 0.0492 (5) | |
| H1A | 0.2133 | 0.4616 | 0.6204 | 0.059* | |
| C2 | 0.18163 (12) | 0.5397 (3) | 0.49388 (14) | 0.0493 (5) | |
| H2 | 0.1239 | 0.5677 | 0.4846 | 0.059* | |
| C3 | 0.21285 (11) | 0.5637 (3) | 0.42623 (12) | 0.0405 (4) | |
| H3 | 0.1766 | 0.6069 | 0.3713 | 0.049* | |
| C4 | 0.29908 (10) | 0.5225 (2) | 0.44139 (10) | 0.0313 (4) | |
| C5 | 0.35276 (12) | 0.4531 (2) | 0.52240 (11) | 0.0396 (4) | |
| H5 | 0.4102 | 0.4230 | 0.5318 | 0.048* | |
| C6 | 0.31953 (13) | 0.4296 (3) | 0.58870 (12) | 0.0481 (5) | |
| H6 | 0.3548 | 0.3828 | 0.6431 | 0.058* | |
| C7 | 0.43547 (11) | 0.2757 (2) | 0.36899 (11) | 0.0378 (4) | |
| H7A | 0.4162 | 0.1957 | 0.4041 | 0.045* | |
| H7B | 0.4827 | 0.3439 | 0.4096 | 0.045* | |
| C8 | 0.46828 (10) | 0.1780 (2) | 0.30692 (11) | 0.0332 (4) | |
| C9 | 0.5000 | 0.2653 (3) | 0.2500 | 0.0320 (5) | |
| H9 | 0.5000 | 0.3845 | 0.2500 | 0.038* | |
| C10 | 0.46821 (12) | 0.0009 (3) | 0.30605 (13) | 0.0429 (4) | |
| H10 | 0.4467 | −0.0593 | 0.3434 | 0.052* | |
| C11 | 0.5000 | −0.0871 (4) | 0.2500 | 0.0504 (7) | |
| H11 | 0.5000 | −0.2063 | 0.2500 | 0.061* | |
| S1 | 0.34361 (3) | 0.56500 (6) | 0.35953 (3) | 0.03403 (15) | |
| N1 | 0.36255 (9) | 0.3882 (2) | 0.31843 (10) | 0.0382 (4) | |
| H1 | 0.3187 (13) | 0.339 (3) | 0.2865 (14) | 0.046* | |
| O1 | 0.42667 (8) | 0.64544 (18) | 0.40359 (9) | 0.0459 (3) | |
| O2 | 0.27803 (9) | 0.65172 (19) | 0.28749 (8) | 0.0502 (4) | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| C1 | 0.0510 (11) | 0.0624 (14) | 0.0390 (10) | −0.0120 (10) | 0.0223 (9) | −0.0015 (9) |
| C2 | 0.0371 (10) | 0.0647 (14) | 0.0507 (11) | −0.0021 (9) | 0.0213 (9) | −0.0029 (10) |
| C3 | 0.0355 (9) | 0.0490 (12) | 0.0342 (9) | 0.0017 (8) | 0.0097 (7) | 0.0017 (8) |
| C4 | 0.0330 (8) | 0.0318 (9) | 0.0282 (7) | −0.0021 (6) | 0.0101 (6) | −0.0021 (6) |
| C5 | 0.0362 (9) | 0.0459 (11) | 0.0343 (8) | 0.0004 (7) | 0.0101 (7) | 0.0019 (7) |
| C6 | 0.0526 (11) | 0.0575 (13) | 0.0307 (9) | −0.0038 (9) | 0.0110 (8) | 0.0074 (8) |
| C7 | 0.0364 (9) | 0.0416 (11) | 0.0367 (9) | 0.0013 (7) | 0.0151 (7) | 0.0044 (8) |
| C8 | 0.0264 (7) | 0.0335 (10) | 0.0379 (8) | −0.0002 (6) | 0.0098 (7) | 0.0026 (7) |
| C9 | 0.0316 (11) | 0.0233 (12) | 0.0409 (12) | 0.000 | 0.0131 (10) | 0.000 |
| C10 | 0.0406 (9) | 0.0337 (10) | 0.0530 (11) | −0.0052 (8) | 0.0155 (9) | 0.0091 (9) |
| C11 | 0.0536 (16) | 0.0234 (14) | 0.0696 (19) | 0.000 | 0.0171 (15) | 0.000 |
| S1 | 0.0348 (2) | 0.0367 (3) | 0.0307 (2) | 0.00170 (17) | 0.01223 (17) | 0.00359 (17) |
| N1 | 0.0313 (7) | 0.0452 (10) | 0.0358 (8) | 0.0011 (6) | 0.0095 (6) | −0.0079 (7) |
| O1 | 0.0439 (7) | 0.0438 (8) | 0.0516 (7) | −0.0117 (6) | 0.0195 (6) | 0.0002 (6) |
| O2 | 0.0544 (8) | 0.0605 (10) | 0.0357 (7) | 0.0195 (7) | 0.0165 (6) | 0.0149 (6) |
Geometric parameters (Å, º) top | C1—C2 | 1.373 (3) | C7—H7A | 0.9700 |
| C1—C6 | 1.373 (3) | C7—H7B | 0.9700 |
| C1—H1A | 0.9300 | C8—C10 | 1.382 (3) |
| C2—C3 | 1.378 (3) | C8—C9 | 1.389 (2) |
| C2—H2 | 0.9300 | C9—C8i | 1.389 (2) |
| C3—C4 | 1.385 (2) | C9—H9 | 0.9300 |
| C3—H3 | 0.9300 | C10—C11 | 1.381 (2) |
| C4—C5 | 1.390 (2) | C10—H10 | 0.9300 |
| C4—S1 | 1.7596 (17) | C11—C10i | 1.381 (2) |
| C5—C6 | 1.379 (3) | C11—H11 | 0.9300 |
| C5—H5 | 0.9300 | S1—O2 | 1.4314 (12) |
| C6—H6 | 0.9300 | S1—O1 | 1.4315 (13) |
| C7—N1 | 1.467 (2) | S1—N1 | 1.6092 (16) |
| C7—C8 | 1.506 (2) | N1—H1 | 0.81 (2) |
| | | |
| C2—C1—C6 | 120.16 (18) | H7A—C7—H7B | 108.1 |
| C2—C1—H1A | 119.9 | C10—C8—C9 | 118.86 (17) |
| C6—C1—H1A | 119.9 | C10—C8—C7 | 120.91 (16) |
| C1—C2—C3 | 120.73 (18) | C9—C8—C7 | 120.22 (17) |
| C1—C2—H2 | 119.6 | C8i—C9—C8 | 121.3 (2) |
| C3—C2—H2 | 119.6 | C8i—C9—H9 | 119.4 |
| C2—C3—C4 | 118.88 (16) | C8—C9—H9 | 119.4 |
| C2—C3—H3 | 120.6 | C11—C10—C8 | 120.31 (19) |
| C4—C3—H3 | 120.6 | C11—C10—H10 | 119.8 |
| C3—C4—C5 | 120.78 (16) | C8—C10—H10 | 119.8 |
| C3—C4—S1 | 120.29 (13) | C10—C11—C10i | 120.4 (3) |
| C5—C4—S1 | 118.89 (13) | C10—C11—H11 | 119.8 |
| C6—C5—C4 | 119.00 (17) | C10i—C11—H11 | 119.8 |
| C6—C5—H5 | 120.5 | O2—S1—O1 | 119.47 (9) |
| C4—C5—H5 | 120.5 | O2—S1—N1 | 105.84 (8) |
| C1—C6—C5 | 120.41 (17) | O1—S1—N1 | 106.67 (8) |
| C1—C6—H6 | 119.8 | O2—S1—C4 | 107.53 (8) |
| C5—C6—H6 | 119.8 | O1—S1—C4 | 107.08 (8) |
| N1—C7—C8 | 110.62 (13) | N1—S1—C4 | 110.10 (8) |
| N1—C7—H7A | 109.5 | C7—N1—S1 | 121.73 (12) |
| C8—C7—H7A | 109.5 | C7—N1—H1 | 114.8 (15) |
| N1—C7—H7B | 109.5 | S1—N1—H1 | 114.1 (15) |
| C8—C7—H7B | 109.5 | | |
| | | |
| C6—C1—C2—C3 | −1.4 (3) | C7—C8—C10—C11 | −178.78 (13) |
| C1—C2—C3—C4 | −0.4 (3) | C8—C10—C11—C10i | −0.31 (11) |
| C2—C3—C4—C5 | 1.9 (3) | C3—C4—S1—O2 | 3.65 (17) |
| C2—C3—C4—S1 | −175.61 (15) | C5—C4—S1—O2 | −173.94 (14) |
| C3—C4—C5—C6 | −1.6 (3) | C3—C4—S1—O1 | 133.19 (15) |
| S1—C4—C5—C6 | 175.99 (15) | C5—C4—S1—O1 | −44.39 (17) |
| C2—C1—C6—C5 | 1.8 (3) | C3—C4—S1—N1 | −111.21 (15) |
| C4—C5—C6—C1 | −0.3 (3) | C5—C4—S1—N1 | 71.20 (16) |
| N1—C7—C8—C10 | −121.05 (17) | C8—C7—N1—S1 | −150.45 (13) |
| N1—C7—C8—C9 | 59.57 (18) | O2—S1—N1—C7 | 170.85 (14) |
| C10—C8—C9—C8i | −0.30 (11) | O1—S1—N1—C7 | 42.63 (16) |
| C7—C8—C9—C8i | 179.09 (15) | C4—S1—N1—C7 | −73.22 (15) |
| C9—C8—C10—C11 | 0.6 (2) | | |
| Symmetry code: (i) −x+1, y, −z+1/2. |
Hydrogen-bond geometry (Å, º) top | D—H···A | D—H | H···A | D···A | D—H···A |
| N1—H1···O2ii | 0.81 (2) | 2.16 (2) | 2.960 (2) | 166 (2) |
| C5—H5···O1iii | 0.93 | 2.55 | 3.461 (2) | 166 |
| C10—H10···O1iv | 0.93 | 2.57 | 3.379 (3) | 146 |
| Symmetry codes: (ii) −x+1/2, y−1/2, −z+1/2; (iii) −x+1, −y+1, −z+1; (iv) x, y−1, z. |
Experimental details
| Crystal data |
| Chemical formula | C20H20N2O4S2 |
| Mr | 416.50 |
| Crystal system, space group | Monoclinic, C2/c |
| Temperature (K) | 296 |
| a, b, c (Å) | 16.4319 (6), 7.8024 (3), 16.1217 (6) |
| β (°) | 111.543 (2) |
| V (Å3) | 1922.54 (12) |
| Z | 4 |
| Radiation type | Mo Kα |
| µ (mm−1) | 0.31 |
| Crystal size (mm) | 0.15 × 0.11 × 0.08 |
| |
| Data collection |
| Diffractometer | Bruker APEXII CCD
diffractometer |
| Absorption correction | – |
No. of measured, independent and
observed [I > 2σ(I)] reflections | 8653, 2346, 1841 |
| Rint | 0.024 |
| (sin θ/λ)max (Å−1) | 0.667 |
| |
| Refinement |
| R[F2 > 2σ(F2)], wR(F2), S | 0.037, 0.103, 1.07 |
| No. of reflections | 2346 |
| No. of parameters | 131 |
| H-atom treatment | H atoms treated by a mixture of independent and constrained refinement |
| Δρmax, Δρmin (e Å−3) | 0.29, −0.34 |
Hydrogen-bond geometry (Å, º) top | D—H···A | D—H | H···A | D···A | D—H···A |
| N1—H1···O2i | 0.81 (2) | 2.16 (2) | 2.960 (2) | 166 (2) |
| C5—H5···O1ii | 0.93 | 2.55 | 3.461 (2) | 166 |
| C10—H10···O1iii | 0.93 | 2.57 | 3.379 (3) | 146 |
| Symmetry codes: (i) −x+1/2, y−1/2, −z+1/2; (ii) −x+1, −y+1, −z+1; (iii) x, y−1, z. |
Acknowledgements
IUK thanks the Higher Education Commission of Pakistan for its financial support under the project to strengthen the Materials Chemistry Laboratory at GCUL.
References
 Bruker (2007). APEX2 and SAINT. Bruker AXS Inc., Madison, Wisconsin, USA. Google Scholar
Bruker (2007). APEX2 and SAINT. Bruker AXS Inc., Madison, Wisconsin, USA. Google Scholar
 Farrugia, L. J. (1997). J. Appl. Cryst. 30, 565. CrossRef IUCr Journals Google Scholar
Farrugia, L. J. (1997). J. Appl. Cryst. 30, 565. CrossRef IUCr Journals Google Scholar
 Khan, I. U., Sheikh, T. A., Ejaz & Harrison, W. T. A. (2011). Acta Cryst. E67, o2371. Google Scholar
Khan, I. U., Sheikh, T. A., Ejaz & Harrison, W. T. A. (2011). Acta Cryst. E67, o2371. Google Scholar
 Sheldrick, G. M. (2008). Acta Cryst. A64, 112–122. Web of Science CrossRef CAS IUCr Journals Google Scholar
Sheldrick, G. M. (2008). Acta Cryst. A64, 112–122. Web of Science CrossRef CAS IUCr Journals Google Scholar
This is an open-access article distributed under the terms of the Creative Commons Attribution (CC-BY) Licence, which permits unrestricted use, distribution, and reproduction in any medium, provided the original authors and source are cited.
 | CRYSTALLOGRAPHIC
COMMUNICATIONS |
ISSN: 2056-9890
Open

access


![Bruker (2007). APEX2 and SAINT. Bruker AXS Inc., Madison, Wisconsin, USA. [Bruker (2007). APEX2 and SAINT. Bruker AXS Inc., Madison, Wisconsin, USA.]](../../../../../../logos/arrows/e_arr.gif) ); cell SAINT (Bruker, 2007
); cell SAINT (Bruker, 2007![Bruker (2007). APEX2 and SAINT. Bruker AXS Inc., Madison, Wisconsin, USA. [Bruker (2007). APEX2 and SAINT. Bruker AXS Inc., Madison, Wisconsin, USA.]](../../../../../../logos/arrows/e_arr.gif) ); data reduction: SAINT; program(s) used to solve structure: SHELXS97 (Sheldrick, 2008
); data reduction: SAINT; program(s) used to solve structure: SHELXS97 (Sheldrick, 2008![Sheldrick, G. M. (2008). Acta Cryst. A64, 112-122. [Sheldrick, G. M. (2008). Acta Cryst. A64, 112-122.]](../../../../../../logos/arrows/e_arr.gif) ); program(s) used to refine structure: SHELXL97 (Sheldrick, 2008
); program(s) used to refine structure: SHELXL97 (Sheldrick, 2008![Sheldrick, G. M. (2008). Acta Cryst. A64, 112-122. [Sheldrick, G. M. (2008). Acta Cryst. A64, 112-122.]](../../../../../../logos/arrows/e_arr.gif) ); molecular graphics: ORTEP-3 (Farrugia, 1997
); molecular graphics: ORTEP-3 (Farrugia, 1997![Farrugia, L. J. (1997). J. Appl. Cryst. 30, 565. [Farrugia, L. J. (1997). J. Appl. Cryst. 30, 565.]](../../../../../../logos/arrows/e_arr.gif) ); software used to prepare material for publication: SHELXL97.
); software used to prepare material for publication: SHELXL97.![[Figure 1]](./su2323fig1thm.gif)
 Bruker (2007). APEX2 and SAINT. Bruker AXS Inc., Madison, Wisconsin, USA. Google Scholar
Bruker (2007). APEX2 and SAINT. Bruker AXS Inc., Madison, Wisconsin, USA. Google Scholar Farrugia, L. J. (1997). J. Appl. Cryst. 30, 565. CrossRef IUCr Journals Google Scholar
Farrugia, L. J. (1997). J. Appl. Cryst. 30, 565. CrossRef IUCr Journals Google Scholar Khan, I. U., Sheikh, T. A., Ejaz & Harrison, W. T. A. (2011). Acta Cryst. E67, o2371. Google Scholar
Khan, I. U., Sheikh, T. A., Ejaz & Harrison, W. T. A. (2011). Acta Cryst. E67, o2371. Google Scholar Sheldrick, G. M. (2008). Acta Cryst. A64, 112–122. Web of Science CrossRef CAS IUCr Journals Google Scholar
Sheldrick, G. M. (2008). Acta Cryst. A64, 112–122. Web of Science CrossRef CAS IUCr Journals Google Scholar
 journal menu
journal menu






 access
access![[Scheme 1]](su2323scheme1.gif)
![[-x+{\script{1\over 2}}, y-{\script{1\over 2}}, -z+{\script{1\over 2}}]](teximages/su2323fi2.gif) ; (ii)
; (ii)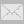 

As part of our ongoing structural studies of sulfonamides (Khan et al., 2011), the synthesis and structure of the title compound are described herein.
The complete molecule of the title compound is generated by crystallographic twofold symmetry (Fig. 1), with atoms C9 and C11 lying on the rotation axis. The diehdral angle between the central benzene ring and the pendant ring is 68.42 (6)°. The dihedral angle between the two pendant rings is 45.11 (5)°. The conformation of the C4—S1—N1—C7 fragment is approximately gauche [-73.22 (15)°] whereas the torsion angle for S1—N1—C8—C9 of -150.45 (13)° indicates a conformation between gauche and anti. The bond-angle sum for N1 of 350.6° seems to indicate a valence state close to sp2 hybridization.
In the crystal, the molecules are linked by N—H···O hydrogen bonds (Table 1), to generate corrugated (010) sheets (Fig. 2). Weak aromatic π-π stacking [centroid-centroid separation = 3.8925 (12) and 3.9777 (12) Å] occurs between the layers and weak C—H···O interactions may help to consolidate the packing.