Buy article online - an online subscription or single-article purchase is required to access this article.
The title compound, C
22H
30N
2O
2·H
2O, is an 18-membered diaza-crown ether ligand containing two ether O and two aza N atoms. In the macrocyclic ring, the mean N

O distance is 4.526 (4) Å. The macrocyclic inner-hole size, estimated as twice the mean distance of the donor atoms from their centroid, is
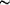
2.29 Å.
Supporting information
CCDC reference: 237948
The title compound, (I), was obtained from the reaction of 1,4-bis(salicyloxy)butane (0.5 g, 1.65 mmol; Hökelek, Kaya & Kılıç, 2001) in methanol (100 ml) and 1,4-diaminobutane (0.26 ml, 2.47 mmol) in methanol (50 ml). Argon was passed over the reaction mixture, and the mixture was refluxed for 1 h, after which excess amounts of borax (2.51 g, 6.60 mmol) and sodium borohydride (0.25 g, 6.60 mmol) were partially added. The reduction was completed after 2 h. The solvent was then evaporated under reduced pressure. The residue was dissolved in chloroform and extracted with water, and the chloroform layers were collected and evaporated under reduced pressure. The oily residue was crystallized from n-heptane (yield 0.58 g, 56%; m.p. 352 K).
Data collection: CAD-4 EXPRESS (Enraf Nonius, 1994); cell refinement: CAD-4 EXPRESS; data reduction: XCAD4 (Harms & Wocadlo, 1995); program(s) used to solve structure: SHELXS97 (Sheldrick, 1997); program(s) used to refine structure: SHELXL97 (Sheldrick, 1997); molecular graphics: ORTEP-3 for Windows (Farrugia, 1997); software used to prepare material for publication: WinGX publication routines (Farrugia, 1999).
1,14-Dioxa-5,10-diaza-2,3:12,13-dibenzocyclooctadeca-2,12-diene monohydrate
top
Crystal data top
| C22H30N2O2·H2O | F(000) = 808 |
| Mr = 372.50 | Dx = 1.183 Mg m−3 |
| Monoclinic, P21/c | Mo Kα radiation, λ = 0.71073 Å |
| Hall symbol: -P 2ybc | Cell parameters from 25 reflections |
| a = 8.6247 (12) Å | θ = 10–18° |
| b = 14.7854 (10) Å | µ = 0.08 mm−1 |
| c = 16.6181 (14) Å | T = 293 K |
| β = 99.352 (12)° | Block, colourless |
| V = 2091.0 (4) Å3 | 0.40 × 0.25 × 0.25 mm |
| Z = 4 | |
Data collection top
Enraf–Nonius CAD-4
diffractometer | Rint = 0.024 |
| Radiation source: fine-focus sealed tube | θmax = 26.3°, θmin = 2.4° |
| Graphite monochromator | h = 0→10 |
| non–profiled ω scans | k = −18→0 |
| 4475 measured reflections | l = −20→20 |
| 4190 independent reflections | 3 standard reflections every 120 min |
| 2237 reflections with I > 2σ(I) | intensity decay: 1% |
Refinement top
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.050 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.158 | H atoms treated by a mixture of independent and constrained refinement |
| S = 1.01 | w = 1/[σ2(Fo2) + (0.0633P)2 + 0.5446P]
where P = (Fo2 + 2Fc2)/3 |
| 4190 reflections | (Δ/σ)max < 0.001 |
| 356 parameters | Δρmax = 0.41 e Å−3 |
| 0 restraints | Δρmin = −0.19 e Å−3 |
Crystal data top
| C22H30N2O2·H2O | V = 2091.0 (4) Å3 |
| Mr = 372.50 | Z = 4 |
| Monoclinic, P21/c | Mo Kα radiation |
| a = 8.6247 (12) Å | µ = 0.08 mm−1 |
| b = 14.7854 (10) Å | T = 293 K |
| c = 16.6181 (14) Å | 0.40 × 0.25 × 0.25 mm |
| β = 99.352 (12)° | |
Data collection top
Enraf–Nonius CAD-4
diffractometer | Rint = 0.024 |
| 4475 measured reflections | 3 standard reflections every 120 min |
| 4190 independent reflections | intensity decay: 1% |
| 2237 reflections with I > 2σ(I) | |
Refinement top
| R[F2 > 2σ(F2)] = 0.050 | 0 restraints |
| wR(F2) = 0.158 | H atoms treated by a mixture of independent and constrained refinement |
| S = 1.01 | Δρmax = 0.41 e Å−3 |
| 4190 reflections | Δρmin = −0.19 e Å−3 |
| 356 parameters | |
Special details top
Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes. |
Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2, conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2 are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top| | x | y | z | Uiso*/Ueq | |
| O | 0.3867 (3) | 0.31760 (17) | 0.14114 (17) | 0.0969 (7) | |
| C1 | 0.2804 (3) | 0.61633 (15) | 0.14932 (14) | 0.0553 (6) | |
| O2 | 0.24524 (18) | 0.63035 (10) | 0.22599 (9) | 0.0602 (4) | |
| C3 | 0.3435 (4) | 0.69184 (19) | 0.27798 (17) | 0.0669 (7) | |
| C4 | 0.2975 (4) | 0.68882 (18) | 0.36109 (17) | 0.0675 (7) | |
| C5 | 0.2965 (4) | 0.59452 (18) | 0.39728 (16) | 0.0638 (7) | |
| C6 | 0.3203 (3) | 0.59607 (18) | 0.48886 (16) | 0.0630 (7) | |
| O7 | 0.2752 (2) | 0.51010 (11) | 0.51630 (10) | 0.0708 (5) | |
| C8 | 0.2822 (3) | 0.49612 (16) | 0.59858 (15) | 0.0576 (6) | |
| C9 | 0.3633 (3) | 0.5513 (2) | 0.65858 (17) | 0.0668 (7) | |
| C10 | 0.3682 (3) | 0.5292 (2) | 0.73964 (18) | 0.0724 (8) | |
| C11 | 0.2921 (4) | 0.4537 (2) | 0.76098 (19) | 0.0761 (8) | |
| C12 | 0.2099 (3) | 0.4007 (2) | 0.70109 (18) | 0.0720 (7) | |
| C13 | 0.2023 (3) | 0.41994 (16) | 0.61892 (15) | 0.0596 (6) | |
| C14 | 0.1114 (3) | 0.3611 (2) | 0.55346 (19) | 0.0713 (7) | |
| N15 | 0.2147 (3) | 0.30138 (15) | 0.51556 (13) | 0.0683 (6) | |
| C16 | 0.1311 (4) | 0.2409 (2) | 0.45236 (17) | 0.0866 (9) | |
| C17 | 0.0518 (4) | 0.2830 (2) | 0.37635 (17) | 0.0840 (9) | |
| C18 | 0.1595 (4) | 0.3358 (2) | 0.32873 (18) | 0.0706 (7) | |
| C19 | 0.0730 (3) | 0.3814 (2) | 0.25617 (18) | 0.0693 (7) | |
| N20 | 0.1778 (2) | 0.43240 (14) | 0.21247 (13) | 0.0594 (5) | |
| C21 | 0.0927 (3) | 0.48672 (19) | 0.14543 (17) | 0.0668 (7) | |
| C22 | 0.2024 (3) | 0.54426 (16) | 0.10655 (14) | 0.0592 (6) | |
| C23 | 0.2339 (4) | 0.5267 (2) | 0.02881 (17) | 0.0775 (9) | |
| C24 | 0.3384 (4) | 0.5790 (2) | −0.0057 (2) | 0.0875 (10) | |
| C25 | 0.4126 (4) | 0.6486 (2) | 0.03730 (18) | 0.0779 (8) | |
| C26 | 0.3862 (3) | 0.66828 (19) | 0.11481 (16) | 0.0659 (7) | |
| H1 | 0.332 (3) | 0.279 (2) | 0.0999 (18) | 0.089 (10)* | |
| H2 | 0.328 (5) | 0.351 (3) | 0.169 (2) | 0.145 (16)* | |
| H15 | 0.286 (4) | 0.338 (2) | 0.4960 (17) | 0.092 (10)* | |
| H16A | 0.2061 | 0.1972 | 0.4382 | 0.104* | |
| H16B | 0.0529 | 0.2076 | 0.4762 | 0.104* | |
| H17A | −0.0287 | 0.3237 | 0.3896 | 0.101* | |
| H17B | −0.0005 | 0.2360 | 0.3413 | 0.101* | |
| H20 | 0.236 (3) | 0.4676 (16) | 0.2468 (14) | 0.064 (8)* | |
| H31 | 0.455 (3) | 0.6725 (17) | 0.2787 (14) | 0.071 (8)* | |
| H32 | 0.325 (3) | 0.7528 (19) | 0.2511 (15) | 0.082 (8)* | |
| H41 | 0.375 (3) | 0.729 (2) | 0.3962 (17) | 0.096 (9)* | |
| H42 | 0.200 (4) | 0.719 (2) | 0.3577 (18) | 0.111 (12)* | |
| H51 | 0.383 (3) | 0.5566 (18) | 0.3796 (16) | 0.086 (8)* | |
| H52 | 0.195 (3) | 0.5659 (18) | 0.3783 (15) | 0.078 (8)* | |
| H61 | 0.261 (3) | 0.6439 (16) | 0.5101 (13) | 0.060 (7)* | |
| H62 | 0.433 (3) | 0.6059 (17) | 0.5135 (14) | 0.075 (8)* | |
| H91 | 0.418 (3) | 0.6011 (19) | 0.6423 (15) | 0.081 (9)* | |
| H101 | 0.426 (3) | 0.5683 (18) | 0.7837 (16) | 0.085 (8)* | |
| H111 | 0.296 (3) | 0.4386 (17) | 0.8163 (16) | 0.072 (8)* | |
| H121 | 0.156 (3) | 0.3486 (17) | 0.7137 (15) | 0.076 (8)* | |
| H141 | 0.038 (3) | 0.3211 (18) | 0.5780 (15) | 0.083 (8)* | |
| H142 | 0.048 (3) | 0.4016 (19) | 0.5102 (17) | 0.094 (9)* | |
| H181 | 0.248 (2) | 0.2955 (13) | 0.3217 (10) | 0.031 (5)* | |
| H182 | 0.213 (3) | 0.3833 (18) | 0.3664 (15) | 0.079 (8)* | |
| H191 | −0.008 (4) | 0.423 (2) | 0.2754 (18) | 0.103 (10)* | |
| H192 | 0.018 (3) | 0.3336 (19) | 0.2150 (16) | 0.089 (8)* | |
| H211 | 0.040 (3) | 0.4423 (17) | 0.1045 (15) | 0.078 (8)* | |
| H212 | 0.007 (3) | 0.5243 (16) | 0.1653 (14) | 0.068 (7)* | |
| H231 | 0.186 (4) | 0.477 (2) | 0.0036 (18) | 0.094 (10)* | |
| H241 | 0.360 (3) | 0.566 (2) | −0.0599 (19) | 0.102 (10)* | |
| H251 | 0.483 (4) | 0.683 (2) | 0.0150 (18) | 0.100 (11)* | |
| H261 | 0.443 (3) | 0.7167 (16) | 0.1457 (14) | 0.065 (7)* | |
Atomic displacement parameters (Å2) top| | U11 | U22 | U33 | U12 | U13 | U23 |
| O | 0.0697 (14) | 0.0952 (16) | 0.1222 (19) | 0.0128 (13) | 0.0053 (14) | −0.0249 (15) |
| C1 | 0.0541 (13) | 0.0553 (14) | 0.0554 (14) | 0.0112 (12) | 0.0057 (11) | 0.0064 (11) |
| O2 | 0.0618 (10) | 0.0579 (9) | 0.0614 (10) | −0.0066 (8) | 0.0113 (8) | −0.0065 (8) |
| C3 | 0.080 (2) | 0.0562 (15) | 0.0659 (17) | −0.0142 (15) | 0.0170 (14) | −0.0059 (13) |
| C4 | 0.080 (2) | 0.0538 (15) | 0.0720 (18) | −0.0115 (15) | 0.0221 (15) | −0.0084 (13) |
| C5 | 0.0676 (17) | 0.0560 (15) | 0.0700 (17) | −0.0017 (14) | 0.0177 (14) | −0.0048 (12) |
| C6 | 0.0601 (16) | 0.0580 (15) | 0.0710 (17) | −0.0070 (14) | 0.0113 (14) | −0.0023 (13) |
| O7 | 0.0838 (13) | 0.0605 (10) | 0.0671 (11) | −0.0139 (9) | 0.0096 (9) | 0.0006 (8) |
| C8 | 0.0518 (14) | 0.0614 (15) | 0.0593 (15) | 0.0056 (12) | 0.0081 (11) | 0.0012 (12) |
| C9 | 0.0609 (16) | 0.0659 (17) | 0.0731 (19) | 0.0007 (14) | 0.0095 (13) | −0.0053 (14) |
| C10 | 0.0660 (18) | 0.081 (2) | 0.0662 (18) | 0.0136 (15) | 0.0003 (14) | −0.0124 (16) |
| C11 | 0.085 (2) | 0.082 (2) | 0.0623 (19) | 0.0187 (17) | 0.0124 (15) | 0.0041 (16) |
| C12 | 0.0736 (18) | 0.0679 (17) | 0.076 (2) | 0.0059 (16) | 0.0173 (15) | 0.0087 (16) |
| C13 | 0.0517 (14) | 0.0587 (14) | 0.0683 (16) | 0.0067 (12) | 0.0094 (12) | 0.0043 (12) |
| C14 | 0.0565 (16) | 0.0691 (17) | 0.086 (2) | −0.0116 (14) | 0.0047 (15) | 0.0068 (16) |
| N15 | 0.0714 (15) | 0.0669 (14) | 0.0667 (14) | −0.0137 (12) | 0.0116 (12) | −0.0034 (11) |
| C16 | 0.104 (2) | 0.0714 (18) | 0.082 (2) | −0.0320 (17) | 0.0081 (17) | 0.0023 (15) |
| C17 | 0.086 (2) | 0.0807 (19) | 0.083 (2) | −0.0352 (16) | 0.0059 (16) | 0.0011 (16) |
| C18 | 0.0724 (19) | 0.0649 (17) | 0.0731 (18) | −0.0212 (16) | 0.0074 (15) | −0.0017 (14) |
| C19 | 0.0672 (17) | 0.0681 (17) | 0.0729 (18) | −0.0138 (15) | 0.0123 (14) | −0.0030 (15) |
| N20 | 0.0591 (13) | 0.0573 (12) | 0.0600 (12) | −0.0072 (10) | 0.0046 (10) | −0.0037 (10) |
| C21 | 0.0633 (17) | 0.0633 (16) | 0.0688 (17) | 0.0003 (14) | −0.0040 (14) | −0.0057 (14) |
| C22 | 0.0596 (15) | 0.0571 (14) | 0.0575 (15) | 0.0112 (12) | −0.0011 (12) | 0.0041 (12) |
| C23 | 0.098 (2) | 0.0717 (19) | 0.0588 (18) | 0.0161 (17) | −0.0004 (16) | −0.0011 (15) |
| C24 | 0.108 (3) | 0.096 (2) | 0.0608 (19) | 0.023 (2) | 0.0191 (18) | 0.0117 (18) |
| C25 | 0.082 (2) | 0.085 (2) | 0.068 (2) | 0.0112 (18) | 0.0170 (16) | 0.0241 (17) |
| C26 | 0.0686 (17) | 0.0646 (16) | 0.0639 (17) | 0.0019 (14) | 0.0086 (14) | 0.0114 (14) |
Geometric parameters (Å, º) top
| O—H2 | 0.89 (4) | C3—H32 | 1.01 (3) |
| O—H1 | 0.96 (3) | C5—C4 | 1.519 (4) |
| O2—C1 | 1.373 (3) | C5—H51 | 1.02 (3) |
| O2—C3 | 1.433 (3) | C5—H52 | 0.98 (3) |
| O7—C8 | 1.375 (3) | C26—C25 | 1.375 (4) |
| O7—C6 | 1.426 (3) | C26—H261 | 0.97 (2) |
| C13—C12 | 1.386 (4) | C9—C10 | 1.380 (4) |
| C13—C8 | 1.390 (3) | C9—H91 | 0.94 (3) |
| C13—C14 | 1.509 (4) | C18—C17 | 1.529 (4) |
| C8—C9 | 1.386 (4) | C18—H181 | 0.991 (19) |
| N15—C14 | 1.467 (3) | C18—H182 | 1.00 (3) |
| N15—C16 | 1.475 (3) | C25—C24 | 1.354 (4) |
| N15—H15 | 0.92 (3) | C25—H251 | 0.92 (3) |
| C1—C26 | 1.386 (3) | C10—C11 | 1.370 (4) |
| C1—C22 | 1.392 (3) | C10—H101 | 1.00 (3) |
| C22—C23 | 1.387 (4) | C23—C24 | 1.382 (5) |
| C22—C21 | 1.495 (4) | C23—H231 | 0.91 (3) |
| N20—C19 | 1.458 (3) | C11—C12 | 1.371 (4) |
| N20—C21 | 1.470 (3) | C11—H111 | 0.94 (2) |
| N20—H20 | 0.87 (2) | C17—C16 | 1.473 (4) |
| C6—C5 | 1.502 (4) | C17—H17A | 0.9700 |
| C6—H62 | 1.00 (3) | C17—H17B | 0.9700 |
| C6—H61 | 0.97 (2) | C12—H121 | 0.94 (3) |
| C21—H212 | 1.02 (2) | C16—H16A | 0.9700 |
| C21—H211 | 1.00 (3) | C16—H16B | 0.9700 |
| C19—C18 | 1.474 (4) | C4—H42 | 0.95 (3) |
| C19—H192 | 1.04 (3) | C4—H41 | 1.00 (3) |
| C19—H191 | 1.02 (3) | C24—H241 | 0.97 (3) |
| C3—C4 | 1.498 (4) | C14—H142 | 1.02 (3) |
| C3—H31 | 1.00 (3) | C14—H141 | 1.00 (3) |
| | | |
| H2—O—H1 | 117 (3) | C1—C26—H261 | 120.0 (14) |
| C1—O2—C3 | 117.21 (19) | C10—C9—C8 | 119.7 (3) |
| C8—O7—C6 | 118.83 (19) | C10—C9—H91 | 122.0 (16) |
| C12—C13—C8 | 117.4 (2) | C8—C9—H91 | 118.2 (16) |
| C12—C13—C14 | 121.8 (3) | C19—C18—C17 | 112.8 (2) |
| C8—C13—C14 | 120.8 (2) | C19—C18—H181 | 118.4 (11) |
| O7—C8—C9 | 124.2 (2) | C17—C18—H181 | 107.4 (11) |
| O7—C8—C13 | 114.9 (2) | C19—C18—H182 | 108.1 (14) |
| C9—C8—C13 | 120.9 (2) | C17—C18—H182 | 106.9 (15) |
| C14—N15—C16 | 114.2 (2) | H181—C18—H182 | 102.1 (18) |
| C14—N15—H15 | 106.4 (18) | C24—C25—C26 | 121.3 (3) |
| C16—N15—H15 | 112.7 (18) | C24—C25—H251 | 119.8 (19) |
| O2—C1—C26 | 124.1 (2) | C26—C25—H251 | 119 (2) |
| O2—C1—C22 | 115.3 (2) | C11—C10—C9 | 120.3 (3) |
| C26—C1—C22 | 120.6 (2) | C11—C10—H101 | 118.8 (15) |
| C23—C22—C1 | 117.9 (3) | C9—C10—H101 | 120.8 (16) |
| C23—C22—C21 | 122.0 (3) | C24—C23—C22 | 121.5 (3) |
| C1—C22—C21 | 120.2 (2) | C24—C23—H231 | 122.3 (19) |
| C19—N20—C21 | 112.7 (2) | C22—C23—H231 | 116.1 (19) |
| C19—N20—H20 | 108.9 (16) | C10—C11—C12 | 119.4 (3) |
| C21—N20—H20 | 109.6 (16) | C10—C11—H111 | 120.2 (16) |
| O7—C6—C5 | 108.1 (2) | C12—C11—H111 | 120.3 (16) |
| O7—C6—H62 | 107.2 (15) | C16—C17—C18 | 115.0 (3) |
| C5—C6—H62 | 112.4 (14) | C16—C17—H17A | 108.5 |
| O7—C6—H61 | 110.3 (13) | C18—C17—H17A | 108.5 |
| C5—C6—H61 | 112.7 (13) | C16—C17—H17B | 108.5 |
| H62—C6—H61 | 106.1 (19) | C18—C17—H17B | 108.5 |
| N20—C21—C22 | 111.4 (2) | H17A—C17—H17B | 107.5 |
| N20—C21—H212 | 110.5 (14) | C11—C12—C13 | 122.3 (3) |
| C22—C21—H212 | 111.8 (13) | C11—C12—H121 | 121.5 (16) |
| N20—C21—H211 | 105.8 (14) | C13—C12—H121 | 116.3 (16) |
| C22—C21—H211 | 109.1 (14) | C17—C16—N15 | 117.4 (2) |
| H212—C21—H211 | 108 (2) | C17—C16—H16A | 108.0 |
| N20—C19—C18 | 111.8 (2) | N15—C16—H16A | 108.0 |
| N20—C19—H192 | 105.9 (15) | C17—C16—H16B | 108.0 |
| C18—C19—H192 | 109.9 (15) | N15—C16—H16B | 108.0 |
| N20—C19—H191 | 111.2 (17) | H16A—C16—H16B | 107.2 |
| C18—C19—H191 | 107.5 (17) | C3—C4—C5 | 114.3 (2) |
| H192—C19—H191 | 111 (2) | C3—C4—H42 | 107.4 (19) |
| O2—C3—C4 | 108.5 (2) | C5—C4—H42 | 113 (2) |
| O2—C3—H31 | 107.4 (14) | C3—C4—H41 | 105.4 (16) |
| C4—C3—H31 | 112.1 (14) | C5—C4—H41 | 110.8 (16) |
| O2—C3—H32 | 105.3 (15) | H42—C4—H41 | 105 (2) |
| C4—C3—H32 | 112.8 (15) | C25—C24—C23 | 119.4 (3) |
| H31—C3—H32 | 110 (2) | C25—C24—H241 | 119.7 (18) |
| C6—C5—C4 | 112.2 (2) | C23—C24—H241 | 120.9 (18) |
| C6—C5—H51 | 108.4 (15) | N15—C14—C13 | 112.1 (2) |
| C4—C5—H51 | 109.9 (15) | N15—C14—H142 | 110.1 (16) |
| C6—C5—H52 | 107.6 (15) | C13—C14—H142 | 108.9 (16) |
| C4—C5—H52 | 109.4 (15) | N15—C14—H141 | 106.8 (15) |
| H51—C5—H52 | 109 (2) | C13—C14—H141 | 109.6 (15) |
| C25—C26—C1 | 119.4 (3) | H142—C14—H141 | 109 (2) |
| C25—C26—H261 | 120.6 (14) | | |
| | | |
| C6—O7—C8—C9 | −15.6 (4) | O7—C8—C9—C10 | −177.4 (2) |
| C6—O7—C8—C13 | 165.3 (2) | C13—C8—C9—C10 | 1.6 (4) |
| C12—C13—C8—O7 | 177.8 (2) | N20—C19—C18—C17 | 179.2 (2) |
| C14—C13—C8—O7 | −2.2 (3) | C1—C26—C25—C24 | −0.6 (4) |
| C12—C13—C8—C9 | −1.3 (3) | C8—C9—C10—C11 | −0.6 (4) |
| C14—C13—C8—C9 | 178.7 (2) | C1—C22—C23—C24 | −0.3 (4) |
| C3—O2—C1—C26 | −11.2 (3) | C21—C22—C23—C24 | −178.4 (3) |
| C3—O2—C1—C22 | 168.4 (2) | C9—C10—C11—C12 | −0.6 (4) |
| O2—C1—C22—C23 | −179.8 (2) | C19—C18—C17—C16 | −176.2 (3) |
| C26—C1—C22—C23 | −0.1 (3) | C10—C11—C12—C13 | 0.9 (4) |
| O2—C1—C22—C21 | −1.7 (3) | C8—C13—C12—C11 | 0.1 (4) |
| C26—C1—C22—C21 | 177.9 (2) | C14—C13—C12—C11 | −180.0 (3) |
| C8—O7—C6—C5 | −178.0 (2) | C18—C17—C16—N15 | 59.9 (4) |
| C19—N20—C21—C22 | 175.0 (2) | C14—N15—C16—C17 | 67.9 (4) |
| C23—C22—C21—N20 | 109.3 (3) | O2—C3—C4—C5 | 53.4 (4) |
| C1—C22—C21—N20 | −68.7 (3) | C6—C5—C4—C3 | 155.0 (3) |
| C21—N20—C19—C18 | −173.5 (2) | C26—C25—C24—C23 | 0.1 (5) |
| C1—O2—C3—C4 | −173.3 (2) | C22—C23—C24—C25 | 0.4 (4) |
| O7—C6—C5—C4 | 164.3 (2) | C16—N15—C14—C13 | 178.7 (2) |
| O2—C1—C26—C25 | −179.8 (2) | C12—C13—C14—N15 | −102.2 (3) |
| C22—C1—C26—C25 | 0.6 (4) | C8—C13—C14—N15 | 77.8 (3) |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| O—H1···N15i | 0.96 (3) | 1.98 (3) | 2.940 (3) | 179 (3) |
| O—H2···N20 | 0.89 (4) | 1.99 (4) | 2.868 (3) | 170 (4) |
| N15—H15···O7 | 0.92 (3) | 2.57 (3) | 3.130 (3) | 120 (2) |
| N20—H20···O2 | 0.87 (2) | 2.43 (2) | 2.985 (3) | 122 (2) |
| Symmetry code: (i) x, −y+1/2, z−1/2. |
Experimental details
| Crystal data |
| Chemical formula | C22H30N2O2·H2O |
| Mr | 372.50 |
| Crystal system, space group | Monoclinic, P21/c |
| Temperature (K) | 293 |
| a, b, c (Å) | 8.6247 (12), 14.7854 (10), 16.6181 (14) |
| β (°) | 99.352 (12) |
| V (Å3) | 2091.0 (4) |
| Z | 4 |
| Radiation type | Mo Kα |
| µ (mm−1) | 0.08 |
| Crystal size (mm) | 0.40 × 0.25 × 0.25 |
| |
| Data collection |
| Diffractometer | Enraf–Nonius CAD-4
diffractometer |
| Absorption correction | – |
No. of measured, independent and
observed [I > 2σ(I)] reflections | 4475, 4190, 2237 |
| Rint | 0.024 |
| (sin θ/λ)max (Å−1) | 0.623 |
| |
| Refinement |
| R[F2 > 2σ(F2)], wR(F2), S | 0.050, 0.158, 1.01 |
| No. of reflections | 4190 |
| No. of parameters | 356 |
| H-atom treatment | H atoms treated by a mixture of independent and constrained refinement |
| Δρmax, Δρmin (e Å−3) | 0.41, −0.19 |
Selected torsion angles (º) top| C6—O7—C8—C13 | 165.3 (2) | O7—C6—C5—C4 | 164.3 (2) |
| C14—C13—C8—O7 | −2.2 (3) | N20—C19—C18—C17 | 179.2 (2) |
| C3—O2—C1—C22 | 168.4 (2) | C19—C18—C17—C16 | −176.2 (3) |
| O2—C1—C22—C21 | −1.7 (3) | C18—C17—C16—N15 | 59.9 (4) |
| C8—O7—C6—C5 | −178.0 (2) | C14—N15—C16—C17 | 67.9 (4) |
| C19—N20—C21—C22 | 175.0 (2) | O2—C3—C4—C5 | 53.4 (4) |
| C1—C22—C21—N20 | −68.7 (3) | C6—C5—C4—C3 | 155.0 (3) |
| C21—N20—C19—C18 | −173.5 (2) | C16—N15—C14—C13 | 178.7 (2) |
| C1—O2—C3—C4 | −173.3 (2) | C8—C13—C14—N15 | 77.8 (3) |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| O—H1···N15i | 0.96 (3) | 1.98 (3) | 2.940 (3) | 179 (3) |
| O—H2···N20 | 0.89 (4) | 1.99 (4) | 2.868 (3) | 170 (4) |
| N15—H15···O7 | 0.92 (3) | 2.57 (3) | 3.130 (3) | 120 (2) |
| N20—H20···O2 | 0.87 (2) | 2.43 (2) | 2.985 (3) | 122 (2) |
| Symmetry code: (i) x, −y+1/2, z−1/2. |

 Subscribe to Acta Crystallographica Section C: Structural Chemistry
Subscribe to Acta Crystallographica Section C: Structural Chemistry
The full text of this article is available to subscribers to the journal.
If you have already registered and are using a computer listed in your registration details, please email
support@iucr.org for assistance.
 O distance is 4.526 (4) Å. The macrocyclic inner-hole size, estimated as twice the mean distance of the donor atoms from their centroid, is
O distance is 4.526 (4) Å. The macrocyclic inner-hole size, estimated as twice the mean distance of the donor atoms from their centroid, is 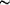 2.29 Å.
2.29 Å.
 Subscribe to Acta Crystallographica Section C: Structural Chemistry
Subscribe to Acta Crystallographica Section C: Structural Chemistry
 journal menu
journal menu








![[Figure 1]](http://journals.iucr.org/c/issues/2004/04/00/ob1170/ob1170fig1thm.gif)
![[Figure 2]](http://journals.iucr.org/c/issues/2004/04/00/ob1170/ob1170fig2thm.gif)

 Subscribe to Acta Crystallographica Section C: Structural Chemistry
Subscribe to Acta Crystallographica Section C: Structural Chemistry



Macrocyclic multidentate ligands, such as the 16-, 17-, 18- and 19-membered ring containing the –OxNy-donor type (where x = 2 and 3, and y = 2 and 3), have been the subject of structural studies as potential metal-ion selective reagents for transition metal-ion recognition (Goodwin et al., 1982; Adam, Leong et al., 1983; Lindoy, 1987; Lindoy et al., 1993; Esteban et al., 2000). Particular metal-ion binding applications (e.g. selective extraction of heavy and precious metals) are of great interest in environmental, inorganic, organic, bio- and coordination chemistry (Esteban et al., 2000; Hökelek, Kaya & Kılıç, 2001; Lindoy, 1997; Hayvalı et al., 1999; Vicente et al., 2000).
Some of the macrocyclic ligands may act as capping ligands, favouring both the blocking of transition metal coordination sites and the formation of discrete metal complexes in inorganic chemistry (Blake & Schröder, 1990; Blake et al., 2000; Nunes et al., 2003). On the other hand, the azamacrocycles, crown ethers and cryptates also play a part in forming host–guest-type inclusion complexes with neutral polar molecules and onium salts (Byriel et al., 2003; Lehn, 1985; Newcomb et al., 1977).
Many investigations have been devoted to the synthetic, thermodynamic and/or structural properties of selective complex formation of a range of transition metal ions and neutral molecules (Adam, Lindoy et al., 1979; Adam, Clarkson et al., 1994; Fenton et al., 1987). However, there are only a limited number of reports concerning the structures of the free macrocyclic multidentate N2O2 and N2O3 donor-type ligands and neutral molecular complexes (Hökelek et al., 1999a,b, 2000; Hökelek, Akduran et al., 2001a,b; Hökelek, Bilge et al., 2001; Hökelek, Kaya & Kılıç, 2001; Hökelek, Bilge & Kılıç, 2002, 2003). The title compound, (I), may be a potential cation selective reagent for transition metal ions. Its dionium salt may also be a potential anion selective reagent. The structure determination of (I) was carried out in order to estimate the macrocyclic ring hole size.
Fig. 1 shows the molecular structure of (I), with the atomic numbering scheme. The macrocyclic ring comprises two ether O and two N atoms and crystallizes with one water molecule. The ligand cavity plays an important role in metal-ion selectivity. The intramolecular O2···N15 [6.877 (3) Å], O7···N20 [5.113 (4) Å], O2···C14 [6.988 (3) Å], O7···C21 [6.118 (4) Å], C3···C16 [7.611 (4) Å] C5···C18 [4.109 (3) Å] and C13···C22 [8.711 (4) Å] distances may indicate the hole size of the macrocyclic ring. When only the N and O atoms are taken into account, the average value of the four N···O distances in the ring is 4.526 (4) Å [O2···N20 = 2.985 (4) Å and O7···N15 = 3.130 (4) Å]. The deviations from the least-squares plane defined by atoms O2, O7, N15 andN 20 are −0.009 (2) Å (O2), 0.012 (2) Å (O7), −0.021 (2) Å (N15) and 0.017 (2) Å (N20).
The macrocyclic inner-hole size, estimated as twice the mean distance of the donor atoms from their centroid, is approximately 2.29 Å, using the 'modified covalent radii' of the Nsp2 (0.66 Å) and Osp3 (0.76 Å) atoms as in the literature method (Goodwin et al., 1982; Adam, Leong et al., 1983; Drummond et al., 1982). The inner hole size of (I) (2.29 Å), which is an 18-membered macrocycle, can be compared with the 16- (1.57 Å; Hökelek et al., 2000), 17- [1.29 (Hökelek et al., 2003) and 2.08 Å (Hökelek, Kaya & Kılıç, 2001)], 18- [1.63 and 1.87 Å (Hökelek, Akduran et al., 2001a), 2.15 Å (Hökelek, Akduran et al., 2001b), and 2.28 Å (Hökelek, Bilge et al., 2001)] and 19-membered (2.53 Å: Hökelek et al., 1999b) multidentate ligand hole sizes. The 15- and 17-membered rings containing the N2O2 and N3O2-donor type ligands afford the NiII and PdII complexes (Bilge et al., 2004). The title ligand, (I), may also give similar complexes with NiII, PdII and other transition metal cations.
In (I), the intermolecular O—H···N hydrogen bonds (Table 2) between the water molecule and atoms N15 and N20 of the macrocycle, and the intramolecular N—H···O close contacts, seem to affect? the macrocyclic ring conformation and, therefore, the macrocyclic inner-hole size. The conformation of the macrocyclic ring is given by the torsion angles. The optimum values of the torsion angles in the macrocyclic ring must be 180° (anti) or 60° (gauche), and in (I), eleven are anti and five are gauche (Table 1). As can be seen from the packing diagram (Fig. 2), the macrocyclic ligands are elongated approximately parallel to the c axis and are stacked along the a axis. The intermolecular hydrogen bonds between the water molecules and the N atoms of the macrocycles result in the formation of zigzag polymeric chains (supramolecules; Lindoy & Atkinson, 2000) parallel to the c axis.