Buy article online - an online subscription or single-article purchase is required to access this article.
The molecules of the title compound, C10H10O5, are linked by O—H O hydrogen bonds into a linear chain along [20
O hydrogen bonds into a linear chain along [20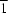 ].
].
 O hydrogen bonds into a linear chain along [20
O hydrogen bonds into a linear chain along [20Supporting information
Crystallographic Information File (CIF) https://doi.org/10.1107/S1600536806027577/ci2104sup1.cif | |
Structure factor file (CIF format) https://doi.org/10.1107/S1600536806027577/ci2104Isup2.hkl |
CCDC reference: 618167
Key indicators
- Single-crystal X-ray study
- T = 295 K
- Mean
![[sigma]](//journals.iucr.org/logos/entities/sigma_rmgif.gif) (C-C) = 0.002 Å
(C-C) = 0.002 Å - R factor = 0.046
- wR factor = 0.153
- Data-to-parameter ratio = 15.2
checkCIF/PLATON results
No syntax errors found
Alert level B PLAT222_ALERT_3_B Large Non-Solvent H Ueq(max)/Ueq(min) ... 4.12 Ratio PLAT731_ALERT_1_B Bond Calc 0.83(5), Rep 0.830(10) ...... 5.00 su-Ra O1 -H1O 1.555 1.555 PLAT735_ALERT_1_B D-H Calc 0.83(5), Rep 0.830(10) ...... 5.00 su-Ra O1 -H1O 1.555 1.555
Alert level C ABSTM02_ALERT_3_C The ratio of expected to reported Tmax/Tmin(RR') is < 0.90 Tmin and Tmax reported: 0.722 0.937 Tmin(prime) and Tmax expected: 0.951 0.991 RR(prime) = 0.803 Please check that your absorption correction is appropriate. PLAT061_ALERT_3_C Tmax/Tmin Range Test RR' too Large ............. 0.80 PLAT062_ALERT_4_C Rescale T(min) & T(max) by ..................... 1.06 PLAT150_ALERT_1_C Volume as Calculated Differs from that Given ... 958.00 Ang-3 PLAT242_ALERT_2_C Check Low Ueq as Compared to Neighbors for C1 PLAT731_ALERT_1_C Bond Calc 0.83(4), Rep 0.840(10) ...... 4.00 su-Ra O5 -H5O 1.555 1.555 PLAT735_ALERT_1_C D-H Calc 0.83(4), Rep 0.840(10) ...... 4.00 su-Ra O5 -H5O 1.555 1.555 PLAT736_ALERT_1_C H...A Calc 1.81(5), Rep 1.82(2) ...... 2.50 su-Ra H1O -O4 1.555 2.664 PLAT736_ALERT_1_C H...A Calc 1.85(4), Rep 1.850(10) ...... 4.00 su-Ra H5O -O2 1.555 2.465
Alert level G ABSTM02_ALERT_3_G When printed, the submitted absorption T values will be replaced by the scaled T values. Since the ratio of scaled T's is identical to the ratio of reported T values, the scaling does not imply a change to the absorption corrections used in the study. Ratio of Tmax expected/reported 1.057 Tmax scaled 0.991 Tmin scaled 0.763
0 ALERT level A = In general: serious problem 3 ALERT level B = Potentially serious problem 9 ALERT level C = Check and explain 1 ALERT level G = General alerts; check 7 ALERT type 1 CIF construction/syntax error, inconsistent or missing data 1 ALERT type 2 Indicator that the structure model may be wrong or deficient 4 ALERT type 3 Indicator that the structure quality may be low 1 ALERT type 4 Improvement, methodology, query or suggestion 0 ALERT type 5 Informative message, check
Computing details top
Data collection: RAPID-AUTO (Rigaku, 1998); cell refinement: RAPID-AUTO; data reduction: CrystalStructure (Rigaku/MSC, 2002); program(s) used to solve structure: SHELXS97 (Sheldrick, 1997); program(s) used to refine structure: SHELXL97 (Sheldrick, 1997); molecular graphics: ORTEPII (Johnson, 1976); software used to prepare material for publication: SHELXL97.
3-(4-Carboxyphenoxy)propionic acid top
Crystal data top
| C10H10O5 | F(000) = 440 |
| Mr = 210.18 | Dx = 1.457 Mg m−3 |
| Monoclinic, P21/c | Mo Kα radiation, λ = 0.71073 Å |
| Hall symbol: -P 2ybc | Cell parameters from 5947 reflections |
| a = 7.813 (4) Å | θ = 3.0–27.5° |
| b = 9.032 (6) Å | µ = 0.12 mm−1 |
| c = 13.631 (9) Å | T = 295 K |
| β = 94.92 (2)° | Plate, colourless |
| V = 958 (1) Å3 | 0.42 × 0.17 × 0.08 mm |
| Z = 4 |
Data collection top
| Rigaku R-AXIS RAPID IP diffractometer | 2188 independent reflections |
| Radiation source: fine-focus sealed tube | 1359 reflections with I > 2σ(I) |
| Graphite monochromator | Rint = 0.035 |
| ω scans | θmax = 27.5°, θmin = 3.0° |
| Absorption correction: multi-scan (ABSCOR; Higashi, 1995) | h = −10→8 |
| Tmin = 0.722, Tmax = 0.937 | k = −11→11 |
| 9176 measured reflections | l = −17→17 |
Refinement top
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.046 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.153 | H atoms treated by a mixture of independent and constrained refinement |
| S = 1.00 | w = 1/[σ2(Fo2) + (0.0912P)2] where P = (Fo2 + 2Fc2)/3 |
| 2188 reflections | (Δ/σ)max = 0.001 |
| 144 parameters | Δρmax = 0.23 e Å−3 |
| 2 restraints | Δρmin = −0.20 e Å−3 |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2) top
| x | y | z | Uiso*/Ueq | ||
| O1 | 0.7570 (2) | 0.4302 (2) | 0.2068 (1) | 0.0650 (5) | |
| O2 | 0.5369 (2) | 0.52999 (2) | 0.1178 (1) | 0.0577 (4) | |
| O3 | 0.3768 (2) | 0.4912 (2) | 0.3441 (1) | 0.0454 (4) | |
| O4 | −0.0388 (2) | 0.9064 (2) | 0.6092 (1) | 0.0628 (5) | |
| O5 | −0.2541 (2) | 0.8132 (2) | 0.5119 (1) | 0.0608 (5) | |
| C1 | 0.5972 (2) | 0.4410 (2) | 0.1810 (1) | 0.0402 (4) | |
| C2 | 0.4826 (2) | 0.3328 (2) | 0.2264 (2) | 0.0461 (5) | |
| C3 | 0.3230 (2) | 0.4016 (2) | 0.2608 (2) | 0.0432 (5) | |
| C4 | 0.2544 (2) | 0.5667 (2) | 0.3894 (1) | 0.0380 (4) | |
| C5 | 0.0802 (2) | 0.5598 (2) | 0.3607 (1) | 0.0426 (5) | |
| C6 | −0.0322 (2) | 0.6431 (2) | 0.4101 (1) | 0.0434 (5) | |
| C7 | 0.0258 (2) | 0.7344 (2) | 0.4873 (1) | 0.0392 (4) | |
| C8 | 0.2008 (2) | 0.7377 (2) | 0.5162 (1) | 0.0441 (5) | |
| C9 | 0.3143 (2) | 0.6542 (2) | 0.4687 (1) | 0.0441 (5) | |
| C10 | −0.0943 (2) | 0.8237 (2) | 0.5398 (1) | 0.0432 (5) | |
| H1O | 0.804 (7) | 0.494 (4) | 0.174 (4) | 0.21 (2)* | |
| H5O | −0.325 (4) | 0.856 (5) | 0.544 (3) | 0.18 (2)* | |
| H2A | 0.5466 | 0.2857 | 0.2820 | 0.055* | |
| H2B | 0.4493 | 0.2564 | 0.1785 | 0.055* | |
| H3A | 0.2656 | 0.4617 | 0.2089 | 0.052* | |
| H3B | 0.2442 | 0.3255 | 0.2790 | 0.052* | |
| H5 | 0.0396 | 0.4995 | 0.3086 | 0.051* | |
| H6 | −0.1493 | 0.6379 | 0.3913 | 0.052* | |
| H8 | 0.2414 | 0.7976 | 0.5686 | 0.053* | |
| H9 | 0.4308 | 0.6560 | 0.4894 | 0.053* |
Atomic displacement parameters (Å2) top
| U11 | U22 | U33 | U12 | U13 | U23 | |
| O1 | 0.0463 (8) | 0.0904 (12) | 0.0569 (10) | −0.0101 (8) | −0.0033 (7) | 0.0161 (9) |
| O2 | 0.0430 (8) | 0.0649 (10) | 0.0663 (10) | 0.0041 (7) | 0.0112 (7) | 0.0194 (8) |
| O3 | 0.0385 (7) | 0.0534 (8) | 0.0446 (8) | 0.0016 (6) | 0.0059 (6) | −0.0088 (6) |
| O4 | 0.0530 (9) | 0.0779 (11) | 0.0566 (10) | 0.0052 (8) | 0.0004 (7) | −0.0228 (8) |
| O5 | 0.0387 (8) | 0.0706 (10) | 0.0733 (11) | 0.0035 (7) | 0.0067 (7) | −0.0202 (8) |
| C1 | 0.0406 (9) | 0.0434 (10) | 0.0368 (10) | 0.0016 (8) | 0.0047 (8) | −0.0052 (8) |
| C2 | 0.0492 (11) | 0.0406 (11) | 0.0497 (12) | −0.0001 (9) | 0.0117 (9) | −0.0005 (9) |
| C3 | 0.0409 (10) | 0.0468 (10) | 0.0425 (11) | −0.0033 (8) | 0.0065 (8) | −0.0028 (8) |
| C4 | 0.0360 (9) | 0.0422 (10) | 0.0361 (10) | −0.0008 (8) | 0.0049 (8) | 0.0023 (8) |
| C5 | 0.0373 (9) | 0.0488 (11) | 0.0410 (10) | −0.0017 (8) | −0.0001 (8) | −0.0058 (9) |
| C6 | 0.0330 (9) | 0.0523 (11) | 0.0446 (11) | 0.0001 (8) | 0.0010 (8) | 0.0005 (9) |
| C7 | 0.0374 (10) | 0.0423 (10) | 0.0384 (10) | 0.0004 (8) | 0.0057 (8) | 0.0017 (8) |
| C8 | 0.0401 (10) | 0.0500 (11) | 0.0420 (11) | −0.0022 (8) | 0.0011 (8) | −0.0060 (9) |
| C9 | 0.0336 (9) | 0.0525 (11) | 0.0455 (11) | −0.0014 (8) | −0.0003 (8) | −0.0027 (9) |
| C10 | 0.0390 (10) | 0.0495 (11) | 0.0413 (10) | −0.0007 (9) | 0.0037 (9) | 0.0016 (9) |
Geometric parameters (Å, º) top
| O1—C1 | 1.271 (2) | C7—C10 | 1.469 (3) |
| O2—C1 | 1.241 (2) | C8—C9 | 1.369 (3) |
| O3—C4 | 1.364 (2) | O1—H1O | 0.83 (1) |
| O3—C3 | 1.428 (2) | O5—H5O | 0.84 (1) |
| O4—C10 | 1.253 (2) | C2—H2A | 0.97 |
| O5—C10 | 1.277 (2) | C2—H2B | 0.97 |
| C1—C2 | 1.494 (3) | C3—H3A | 0.97 |
| C2—C3 | 1.504 (3) | C3—H3B | 0.97 |
| C4—C5 | 1.385 (2) | C5—H5 | 0.93 |
| C4—C9 | 1.387 (3) | C6—H6 | 0.93 |
| C5—C6 | 1.375 (3) | C8—H8 | 0.93 |
| C6—C7 | 1.382 (3) | C9—H9 | 0.93 |
| C7—C8 | 1.390 (2) | ||
| C4—O3—C3 | 118.3 (1) | C10—O5—H5O | 119 (3) |
| O2—C1—O1 | 123.1 (2) | C1—C2—H2A | 108.8 |
| O2—C1—C2 | 120.3 (2) | C3—C2—H2A | 108.8 |
| O1—C1—C2 | 116.4 (2) | C1—C2—H2B | 108.8 |
| C1—C2—C3 | 113.7 (2) | C3—C2—H2B | 108.8 |
| O3—C3—C2 | 106.7 (2) | H2A—C2—H2B | 107.7 |
| O3—C4—C5 | 124.0 (2) | O3—C3—H3A | 110.4 |
| O3—C4—C9 | 115.7 (2) | C2—C3—H3A | 110.4 |
| C5—C4—C9 | 120.2 (2) | O3—C3—H3B | 110.4 |
| C6—C5—C4 | 119.4 (2) | C2—C3—H3B | 110.4 |
| C5—C6—C7 | 121.1 (2) | H3A—C3—H3B | 108.6 |
| C6—C7—C8 | 118.6 (2) | C6—C5—H5 | 120.3 |
| C6—C7—C10 | 121.1 (2) | C4—C5—H5 | 120.3 |
| C8—C7—C10 | 120.3 (2) | C5—C6—H6 | 119.4 |
| C9—C8—C7 | 121.1 (2) | C7—C6—H6 | 119.4 |
| C8—C9—C4 | 119.6 (2) | C9—C8—H8 | 119.5 |
| O4—C10—O5 | 122.4 (2) | C7—C8—H8 | 119.5 |
| O4—C10—C7 | 120.0 (2) | C8—C9—H9 | 120.2 |
| O5—C10—C7 | 117.6 (2) | C4—C9—H9 | 120.2 |
| C1—O1—H1O | 106 (4) | ||
| O2—C1—C2—C3 | −48.1 (3) | C5—C6—C7—C10 | 179.8 (2) |
| O1—C1—C2—C3 | 135.7 (2) | C6—C7—C8—C9 | 0.9 (3) |
| C4—O3—C3—C2 | 179.9 (2) | C10—C7—C8—C9 | 179.4 (2) |
| C1—C2—C3—O3 | −69.8 (2) | C7—C8—C9—C4 | 1.1 (3) |
| C3—O3—C4—C5 | 1.3 (3) | O3—C4—C9—C8 | 177.7 (2) |
| C3—O3—C4—C9 | −178.8 (2) | C5—C4—C9—C8 | −2.3 (3) |
| O3—C4—C5—C6 | −178.6 (2) | C6—C7—C10—O4 | −179.6 (2) |
| C9—C4—C5—C6 | 1.5 (3) | C8—C7—C10—O4 | 1.9 (3) |
| C4—C5—C6—C7 | 0.5 (3) | C6—C7—C10—O5 | 0.4 (3) |
| C5—C6—C7—C8 | −1.7 (3) | C8—C7—C10—O5 | −178.1 (2) |
Hydrogen-bond geometry (Å, º) top
| D—H···A | D—H | H···A | D···A | D—H···A |
| O1—H1O···O4i | 0.83 (1) | 1.82 (2) | 2.618 (2) | 162 (5) |
| O5—H5O···O2ii | 0.84 (1) | 1.85 (1) | 2.676 (2) | 173 (5) |
| Symmetry codes: (i) x+1, −y+3/2, z−1/2; (ii) x−1, −y+3/2, z+1/2. |

 Subscribe to Acta Crystallographica Section E: Crystallographic Communications
Subscribe to Acta Crystallographica Section E: Crystallographic Communications
The full text of this article is available to subscribers to the journal.
- Information on subscribing
- Sample issue
- If you have already subscribed, you may need to register

 journal menu
journal menu











 Alert level B
Alert level B
 Alert level C
Alert level C
 Alert level G
Alert level G



